(→Proof of protein production) |
(→Proof of swimming recovery) |
||
| Line 38: | Line 38: | ||
===Proof of swimming recovery=== | ===Proof of swimming recovery=== | ||
| − | We have made a biobrick BbaK1951008 ables to produce Flagellin (FliC protein of the flagellum). In the aim to test the flagellin integrity, we made a fliC mutant in a <i>E.coli W3110</i> strain by transduction using phage P1 (protocol available on our website). | + | We have made a biobrick BbaK1951008 ables to produce Flagellin (FliC protein of the flagellum). In the aim to test the flagellin integrity, we made a fliC mutant in a <i>E.coli W3110</i> strain by transduction using phage P1 (protocol available on our website). ==Demonstration of motility complementation== |
| − | [[File:T--Aix-Marseille-- | + | |
| + | In the figure the deletion mutant (lower left sector) shows no swimming motility as expected and a small white colony. | ||
| + | In contrast the wild-type colony (lower right) has a diffuse halo due to swimming cells around the central white colony. | ||
| + | Finally the complemented strain, the deletion mutant complemented with our biobrick, (top panel) shows two colonies with intense | ||
| + | halos surrounding them. | ||
| + | This illustrated clearly that our biobrick can restore motility and is functional. | ||
| + | The intensity of the halo suggests that a greater proportion of the cells are mobile or swimming is in someway better than the wild-type. | ||
| + | |||
| + | |||
| + | [[File:T--Aix-Marseille--result3.jpeg|770px|center|thumb|We investigated if swimming motility was recovered by a knockout FliC strain. | ||
| + | To test complementation with our biobrick strains were stabbed into soft (<b>Claire check</b> 0.5%) LB agar plates, | ||
| + | and incubated at 37°C for 4 hours. | ||
| + | Three strains are shown: | ||
| + | <i> Escherichia coli</i> W3110 wild-type strain, which has a good swimming capacity (lower right); | ||
| + | a fliC deletion mutant of W3110 (lower left); | ||
| + | and the fliC mutant complemented with BBa_K1951008(top). | ||
| + | ]] | ||
===Microscopie of the flagellum=== | ===Microscopie of the flagellum=== | ||
Result
Our lab experience enabled us to create the final Biobrocks of our project which are a siderophore Desferrioxamine B producer and a flagellin producer.
Mobilisation result
Proof of protein production
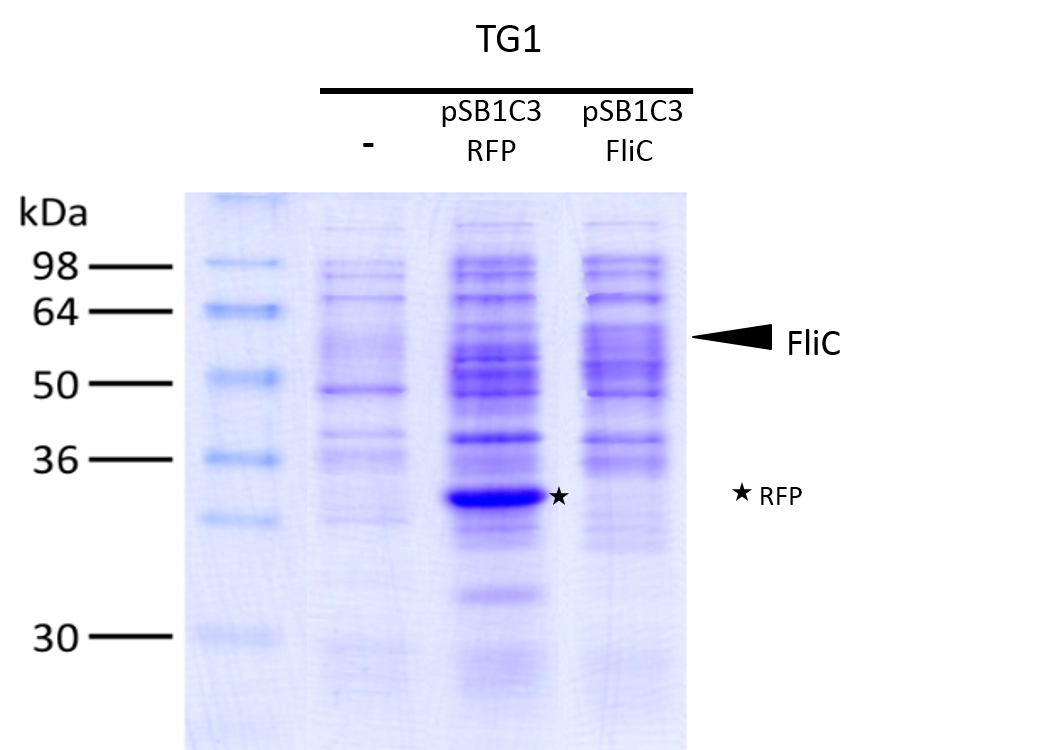
We investigated if the FliC protein was well produced by our biobrick using SDS PAGE.
To to do this we performed SDS PAGE and stained with coomassie blue using cells containing this biobrick in plasmid backbone SDS page and coomassie blue.
From an over night starter, cells were diluted and grown from Abs(600nm)=0.2 to Abs(600nm)=0.6.
Then 1UOD of cells (1.67ml at 0.6OD) was collected and centrifuged at 5000g for 5min.
After removal of the supernatant, the cell pellet was resuspended in 50µL SDS-PAGE sample buffer.
SDS page and coomassie blue The mixture was loaded onto a polyacrylamide gel and migrated during 50min at 180V.
Staining was done using coomassie blue.
The FliC is at mass 51,3kDa, and can be clearly seen in the gel photograph.
Proof of fonctionnality
We investigated if DesA (Lysine decarboxylase) was fonctionnal, by measurement of cadaverine using HPLC with C18 column and proofed our biobrick is a producer of this protein and makes it fonctionnal.

In the futur
We plan to test the whole pathway viability in the first hand. In a second hand, the potential adsorption of platium by our siderophore would be investigate in a rich metal medium.
Biosorption result
Proof of swimming recovery
We have made a biobrick BbaK1951008 ables to produce Flagellin (FliC protein of the flagellum). In the aim to test the flagellin integrity, we made a fliC mutant in a E.coli W3110 strain by transduction using phage P1 (protocol available on our website). ==Demonstration of motility complementation==
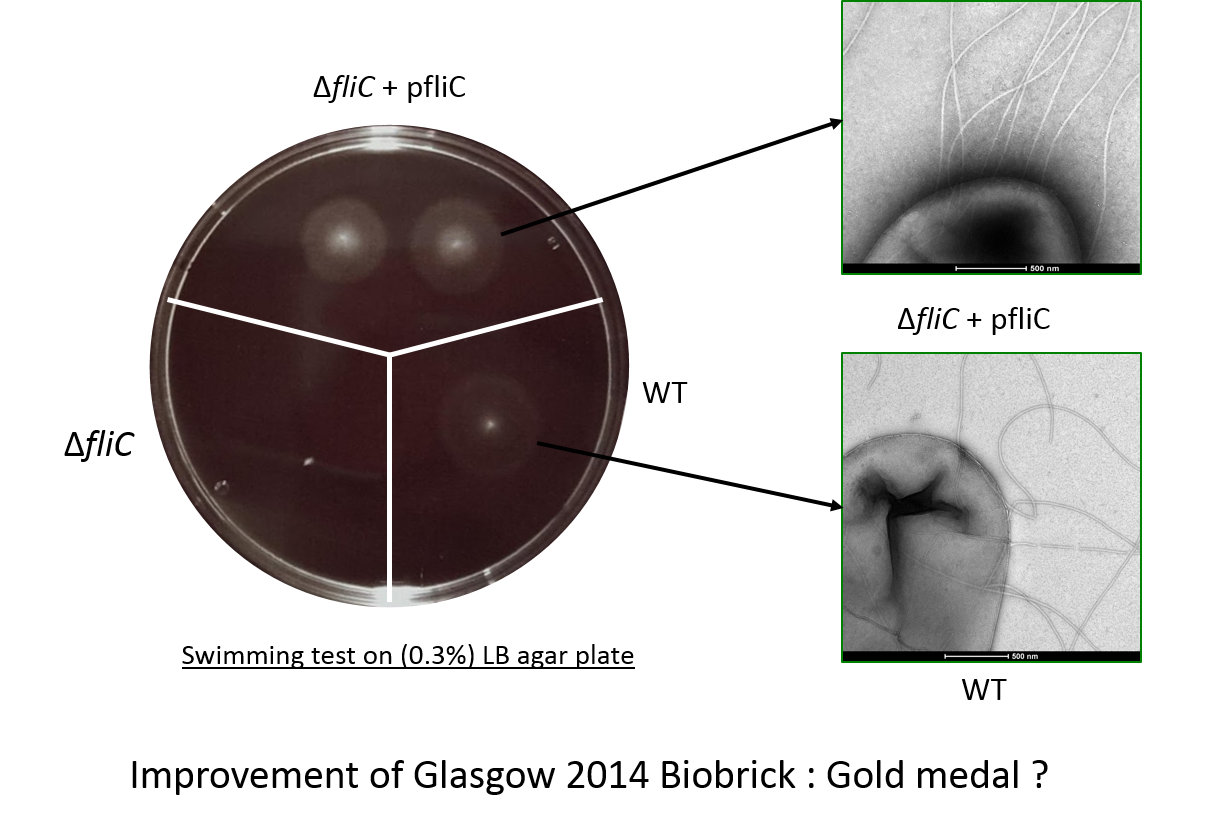
In the figure the deletion mutant (lower left sector) shows no swimming motility as expected and a small white colony.
In contrast the wild-type colony (lower right) has a diffuse halo due to swimming cells around the central white colony.
Finally the complemented strain, the deletion mutant complemented with our biobrick, (top panel) shows two colonies with intense
halos surrounding them.
This illustrated clearly that our biobrick can restore motility and is functional.
The intensity of the halo suggests that a greater proportion of the cells are mobile or swimming is in someway better than the wild-type.
Microscopie of the flagellum
We analysed the fliC mutant complemented by BbaK1951008 using electronic microscopy. This tools allowed us to observe the flagellum integrity recovered and to obtain phenotypic image of our work.
In the futur
We plan to make a proof of concept of the platinum absorption to the surface of the flagellin. In a second time, we want to finish the insertion of the restriction site to allow the cassette insertion ( peptide absorbing specifically precious metal) and test their affinity.
Improvement of FliC E. coli BBa_K342000
This biobrick has been improoved from a previous one designed by Glasgow 2014 team. Please find the link of this biobrick below : http://parts.igem.org/Part:BBa_K1463601
Instead of promotors Bba_J23106 and Bba_J23116, we used strong promoter, strong RBS combination for high expression levels of the flagellin. By the combination of Bba_K880005 and Bba_K1951005, we made a high flagellin expression vector able to recover swimming.
Project achievement in a table
Futur plan for our project
Now that every biobricks have been created and are available, make proofs of concept could allow to envisage investigation about possibilities of an industrial application.
The biologic production of high and specific adsorber could reduce environnemental impact of the metal production and its cost while renew essential precious metals like platinum, for a sustainable production.