Igem2016cjw (Talk | contribs) |
Igem2016cjw (Talk | contribs) |
||
| Line 148: | Line 148: | ||
<div class="col-md-7"> | <div class="col-md-7"> | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
| − | <img src="https://static.igem.org/mediawiki/2016/ | + | <img src="https://static.igem.org/mediawiki/2016/c/c4/IGEM-6803-E4.png" alt="desktop"> |
| − | <p style="font-size:15px"> Fig. | + | <p style="font-size:15px"> Fig.4. Overlap PCR</p> |
</div> | </div> | ||
Revision as of 10:32, 4 October 2016
Experiment of A Controllable Lipid Producer
Overview
Cyanobacteria are excellent organisms for biofuel production. We thus have selected Cyanobacterium Synechocystis sp. PCC 6803 as the source of carbon in our mixed bacteria system. Our target is simply to make the cyanobacteria lyse at the appropriate time by transforming a plasmid contained three bacteriophage-derived lysis genes which were placed downstream of a nickel-inducible signal transduction system into the Synechocystis 6803.

Fig.1. Lipid contents
1. Lipid Producer
Photosynthetic microorganisms, including eukaryotic algae and cyanobacteria, are being optimized to overproduce numerous biofuel. According to previous data, algae accumulate large quantities of lipid as storage materials, but they do this when under stress and growing slowly. By contrast, cyanobacteria accumulate lipids in thylakoid membranes, which are associated with high levels of photosynthesis and a rapid growth rate. Thus, photo-synthetic bacteria have a natural advantage for producing lipids at a high rate. Furthermore, being prokaryotes can be improved by genetic manipulations much more readily than can eukaryotic algae. (Espaux L et al. 2015) Therefore, we decided to do something to make cyanobacteria ,the lipid producer, more appropriate for our project.
Synechococcus elongatus PCC7942 has larger capacity of lipid production than Synechocystis sp. PCC6803 but accumulates most of the product in the cell because of the imbalance of the rates of lipid production and secretion. Initially, we intended to do something to increase lipid secretion by knocking the wzt gene, however, Synechococcus elongatus PCC7942 wasn’t able to revive in two-week shaking cultivation. So we turned into Synechocystis sp.PCC 6803.
2.Lipid Recovery From Biomass
The first goal of our research was to facilitate lipid recovery from biomass. The scientific community widespread disrupts the cyanobacterial cell envelope to achieve the goal. (Seog JL et al. 1998)However, all these methods are not economical for large amounts of biomass or add additional cost and reduce the overall utility of the process. Our target is simply to make the cyanobacteria lyse at the appropriate time. We found that the cyanobacterial cell envelope is composed of 4 layers: the external surface layers ;the outer membrane; the polypeptidoglycan which is considerably thick, and the cytoplasmic membrane.( Hoiczyk E et al. 2000)To break up the peptidoglycan layer, we applied the holin-endolysin lysis strategy used by bacteriophages to exit bacterial cells(Wang IN et al. 2000). Endolysins are peptidoglycan-degrading enzymes that attack the covalent linkages of the peptidoglycans that maintain the integrity of the cell wall. In addition to endolysins, some auxiliary lysis factors are involved in cleaving the oligopeptide linkages between the peptidoglycan and the outer membrane lipoprotein. Holins are small membrane proteins that produce nonspecific lesions (holes) in the

Fig.2. The functions of holin and endolysins in degrading the cell wall
cytoplasmic membrane from within, allow the endolysins and auxiliary lysis factors to gain access to the polypeptidoglycan layers, and trigger the lysis process. In this way, the cell wall is easy to break up.
3. Control The Lysis System
To control the appropriate time, a nickel sensing/responding signal system(Garcia-Dominguez M et al. 2000) was used to control the timing of the expression of phage lysis genes in Synechocystis 6803.
Our strategy for achieving our target is to construct a expression vector pCPC3031-Ni-13-19-15 introduced the Salmonella phage P22 lysis cassette (13-19-15) with a Ampicillin selection marker downstream of the promoter Pni, a nickel responding signal operon. Synechocystis 6803 with the pCPC3031-Ni-13-19-15 will lyse after Ni2+ addition.
Aim
In this part of our project, Cyanobacterium Synechocystis sp. PCC 6803 was selected as a model organism as the source of carbon in our mixed bacteria system. We simply to establish a cell wall disruption process which could make the cyanobacteria lyse at the appropriate time.
Strategies
1. Lysis genes(13-19-15)
P22 gp13,P22 gp19 and P22 gp 15 are holins, endolysins and auxiliary lysis factors respectively. To construct the holin-endolysin lysis system, they should be connected together with defined sequence. First of all, we obtained the three lysis genes which were synthesized by GENEWIZ separately. Then we use PCR to amplify this part. TA cloning and ligation of Blunt-ended DNA on the T vector were our original idea.However, 19 and 15 were spliced via TA cloning according to our presumption. 13 and 19-15 were ligated PCR overlap extension method of Warrens et al.( Warrens AN et al. 1997)

Fig.3. TA cloning and blunt end ligation
2.A Nickel Sensing/Responding Signal System(pCPC3031-Ni)
Ni activates the transcription of downstream genes of Pni and positively autoregulates its own synthesis. The amount of mRNA increased about 20-fold within 4 h after Ni addition. First, Pni was cloned into pCPC3031 and was amplified by using PCR. Then we cut the plasmid with Nru I, after that the lysis genes, 13-19-15, were placed downstream of Pni. What’ more, we handed in the plasimids for sequencing, which confirmed its correctness.
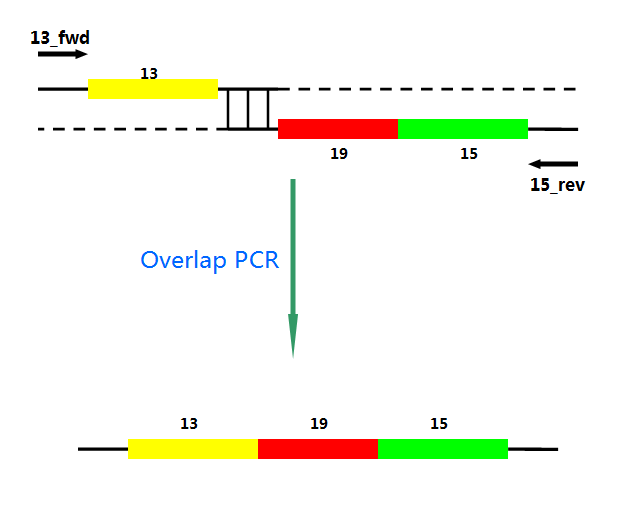
Fig.4. Overlap PCR
6. Verification of ddpX Gene Effect
Just like the verification of RFP mentioned before, the verification of ddpX is carried out in the similar way. The pET21a plasmid is cut by BamH1 and EcoR1 instead of Xba1 and EcoR1 , so that the ddpX can be linked to the cut plasmid pET21a solely. Then we can detect if the cell lysis occurs.
7. Method of Cell Lysis Assay
Cell lysis can be reflected by the OD600 of culture medium. The lower the value of OD600 is than the wild type E.coli at the same condition, the stronger the cell lysis effect will be. The OD600 is detected by 96-well Microplate Reader. In order to know the OD600 value continuously, the detection process works through the time of bacterial growth and we will obtain the OD600-Growing time curve.
8. Chassis selection for TPA Positive Feedback Based Regulation System
As the explanation before, the TPA positive feedback system is derived from the TPA degradation metabolic pathway in Rhodococcus jostii RHA1. Considering the difficulty of conducting gene-scale operation in this unusual organism, we directly synthetize all the gene including tpaK, tpaR, and TILS. At first we want to use E.coli to test this device because of the easy and familiar operation. However, in this situation, we have to transform at least 3 plasmids and this cannot be more difficult for E.coli. Therefore, we use another familiar organism, Saccharomyces cerevisiae, as the chassis. In the preliminary experiment, we successfully transform 3 plasmids into Saccharomyces cerevisiae.
9. Construction of TPA Positive Feedback Based Regulation System
We use common plasmids of Saccharomyces cerevisiae, pRS413, pRS415 and pYES2, to respectively load the TPA transporting protein gene, TPA regulation protein gene and TPA induced RFP gene. First of all, we use PCR to amplify all of these fragments and add different restriction endonuclease cutting sites. Then we cut the plasmids with corresponding restriction endonucleases. Then these cut fragments are linked according to the designed order and transformed into Saccharomyces cerevisiae. We screen for the correctly transformed cell by using the Sc-Ura-Leu-His plate.

Fig.10. The construction process of our TPA Positive Feedback Based regulation system
10. Culture and Expression Condition of Saccharomyces cerevisiae in this experiment
Traditional YPD culture medium (22g/L glucose, 20g/L peptone, 10g/L yeast extracts) is used by us. Sc-Ura-Leu-His culture medium (22g/L glucose, 6.7g/L yeast nitrogen base, 1.224g/L nutrient deficiency mixture without Ura, His, Leu and Trp, 5mg/L Trp) is used to screen for correctly transformed cell. All the cells are cultured in 5mL medium at 30℃ with shaking speed of 200rpm. To induce the expression of RFP, we add TPA with different concentration. We first make up TPA standard solution with TPA concentration of 5g/L. Then we respectively add 0, 1μL, 10μL, 100μL, 1mL standard solution to the culture medium.
Expected Results
PETase and MHETase are two key enzymes in our project. However, as heterologous proteins, the expression of these two enzymes face many problems just like expressing other heterologous proteins before including the formation of inclusion body, the lack of regulation pathway, etc. We design this R-R system in order to express the two enzymes visibly and regularly.
First, we hope to directly observe the expression condition of our enzyme by color, when the inclusion body form, which means the overexpressing, the red color can be observed.
Second, when inclusion body form, the normal way to solve this problem is to use lysozyme and ultrasonic to break the cell and purify the protein, which is complex and time-consuming. We expect the cell lysis will automatically occur when the inclusion body form by using ddpX gene.
Third, we expect the chassis organism can sense the existence of TPA, the hydrolyze product of PET and using TPA as the induction of PETase gene. Thus if the degradation process start, this process can be even enhanced until the PET is used up.
References
[1]Physiologie der Mikroorganismen, Humboldt Universitat zu Berlin, Chausseestr. Misfolded maltose binding protein MalE219 induces the CpxRA envelope stress response by stimulating phosphoryl transfer from CpxA to CpxR. Research in Microbiology 160 (2009) 396-400.
[2]Ivan A. D. Lwssard and Christopher T. Walsh. VanX, a bacterial D-alanyl-D-alanine dipeptidase: Resistance, immunity, or survival function? Proc. Natl. Acad. Sci. USA. Vol. 96, pp. 11028–11032, September, 1999.
[3]Hirofumi Hara, Lindsay D. Eltis, Julian E. Davies. Transcriptomic Analysis Reveals a Bifurcated Terephthalate
Degradation Pathway in Rhodococcus sp. Strain RHA1. Journal of Bacteriology, Mar. 2007, 189(5), 1641–1647.
[4]Molina-Henares, A. J., T. Krell, M. E. Guazzaroni, A. Segura, and J. L. Ramos. 2006. Members of the IclR family of bacterial transcriptional regulators function as activators and/or repressors. FEMS Microbiol. Rev. 30: 157–186.


