Conjugation
We have attempted to conjugate GFP into both G. oxydans and G. hansenii with a Diaminopimelic Acid (DAP) auxotrophic strain of E. coli . The plasmid contains the vector pMMB67EH, the promoter PA-1, GFP and a spectinomycin resistance gene.
The first conjugation was done with KOM strains 4 (G. oxydans) 5 ( G. oxydans and 15 ( L. fermentati ). We attempted these conjugations before sequencing the recipient strains, so that is why we tried to conjugate into L. fermentati . First, a mixture between a KOM strain and the DAP auxotroph strain were plated on a LB+DAP solid medium to allow for conjugation to occur. After 24 hours of incubation, I scraped up the growth and plated each conjugation mixture onto a LB+Spec plate.
Next, we viewed the potential transconjugants on a fluorescence microscope.
We then picked these glowing colonies and then after streaking them out onto more LB+Spec plates, we attempted to use 16s sequencing to confirm successful conjugation. After troubleshooting our 16s procedure, we were finally able to obtain a viable sequencing result. However, all of the glowing colonies were identified as E. coli. For the next round of conjugation, we used a strain of both G. oxydans and Gluconacetobacter hansenii from the American Type Culture Collection (ATCC).
These growths were then scraped up and plated onto a LB+Spec plate.
We then picked isolated colonies and streaked them out onto LB+DAP plates. After using 16s sequencing on the potential transconjugants, we encountered an anomaly. Instead of amplifying the 16s gene, we recieved the sequence of the L,D Transpeptidase gene of E. coli . We plan on repeating the 16s procedure.
Ethanol Reduction
Identifying genes of interest
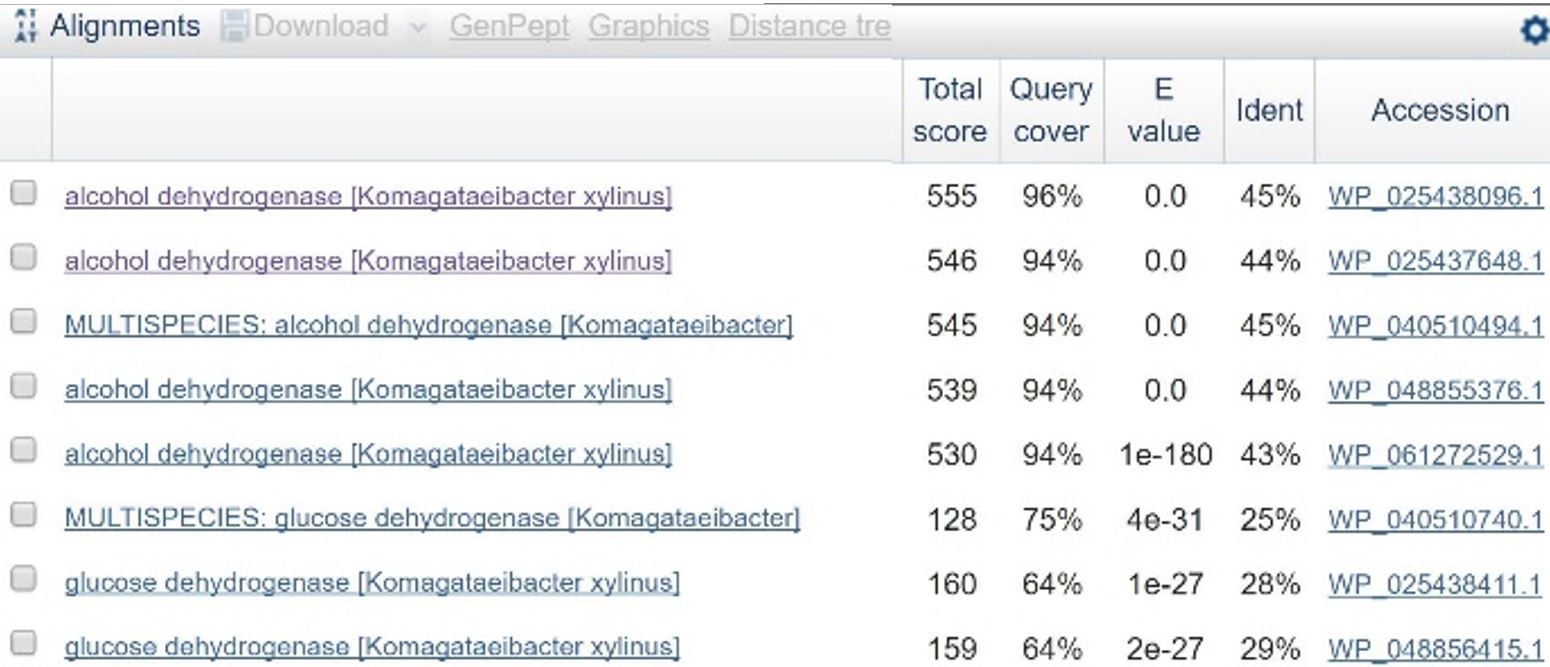
In order to design a construct increasing expression of PQQ-ADH and ALDH in Ga. hansenii, it was necessary to find the genome of the ATCC strain and identify the coding sequences for these genes. The whole genome shotgun sequence for our organism, ATCC 53582, is published on NCBI by J. Abbot (2015) with annotations regarding the functions of specific sequences. Coding sequences are annotated with proposed gene products. Though there are several aldehyde dehydrogenase genes annotated in the genome, there is only one which is described as membrane-bound, matching the description from Mamlouk and Gullo (2013). There are additionally multiple alcohol dehydrogenases. A known amino acid sequence for a homologous PQQ-ADH in Comamonas testosteroni was compared against sequences in the Ga. hansenii genome using BLAST (Table 1). One ADH enzyme found in the Ga. hansenii genome sequence matches the C. testosteroni sequence with a query cover value of 94% and an E value of 0 (third line of table 1).
Creation of Golden Gate parts [[File:T--Austin UTexas--ALDHlinmap.png|thumb|Figure 1: Linear map of the coding sequence for membrane-bound ALDH with a Golden Gate type 3 prefix and suffix. BsmBI and BsaI sites are indicated. The restriction sites at either end are included in the prefix and suffix, but the internal BsaI site must be removed to create a functioning Golden Gate part.|600px]]
In order to assemble the construct, the coding sequences for the genes of interest must be amplified from the Ga. hansenii genome and edited such that they have the correct Golden Gate overhangs and no internal BsaI or BsmBI restriction sites. The sequences were uploaded to Benchling for analysis and planning. The coding sequence for the membrane-bound ALDH contains a BsaI restriction site near the middle of the gene (Figure 1), and the PQQ-ADH coding sequence contains a BsmBI restriction site near the end of the gene (Figure 2). To eliminate the BsaI site in ALDH, primers were designed that would introduce a point mutation at the restriction site. One set of primers, igem2016_KOM_EtOH_01 and igem2016_KOM_EtOH_02, amplifies the sequence upstream of the restriction site, adding a type 3 Golden Gate prefix and removing the restriction site. Another set, igem2016_KOM_EtOH_03 and igem2016_KOM_EtOH_04, amplifies the region downstream of the restriction site, introducing a mutation to the site and adding a type 3 Golden Gate suffix to the end of the gene. These two products will be used in an overlap PCR reaction to create a final product with no BsaI restriction sites and the correct prefix and suffix for assembly. To remove the BsmBI site from the PQQ-ADH coding sequence, a set of primers (igem2016_KOM_EtOH_05 and igem2016_KOM_EtOH_06) was designed to amplify the region upstream of the restriction site and add a Golden Gate type 3 prefix to the beginning of the sequence. The reverse primer additionally adds a mutation to existing BsmBI restriction site and creates a new BsmBI restriction site that will be used to join the piece to a double-stranded DNA, igem2016_KOM_EtOH_07, containing the rest of the gene’s coding sequence appended with a Golden Gate type 3 suffix. The assembly of the PQQ-ADH part will therefore take place in two reactions: one reaction in which the upstream piece of DNA is created, and one reaction in which it is ligated to the gBlock. Table 2 contains more information about each of these oligonucleotides. All were ordered from IDT.
*Will need to insert additional tables and figures. Numbering of tables and figures may need to be adjusted. If anyone is willing to help insert figures and tables in this section, contact me (Stratton) and I'll send everything to be inserted.
| Species | Classification | Brand of Kombucha Isolated From |
|---|---|---|
| Staphylococcus warneri | Bacteria | GT’s Kombucha |
| Staphylococcus epidermidis | Bacteria | GT's Kombucha |
| Gluconobacter oxydans* | Bacteria | GT’s Kombucha |
| Lachancea fermentati* | Yeast | Buddha's Brew |
| Propionibacterium acnes | Bacteria | Buddha's Brew |
| Micrococcus luteus | Bacteria | Buddha's Brew |
| Bacillus pumilus | Bacteria | Buddha's Brew |
| Saccharomyces cerevisiae | Yeast | LIVE Soda Kombucha |
| Schizosaccharomyces pombe* | Yeast | LIVE Soda Kombucha |
(*Indicates a species that is considered vital to the production of kombucha)


pH Sensors
Possessing the ability to monitor the brewing process of kombucha without disturbing the microenvironment and using a very visible color reporter would allow for greater insights as to how the populations of organisms and pH may change due to competition amongst other bacteria and yeast in the beverage and SCOBY of the kombucha. The byproducts produced by the kombucha as it brews causes the tea to become more acidic, leading to our team searching for pH sensitive promoters, and for ways to implement these into kombucha.
Though an acidic sensor was what was required for our kombucha analysis, the identification of sensors in other areas of the pH spectrum were explored as well. Three sequences were identified, the CadC operon for the acidic range, CpxA-CpxR complex for the neutral range, and the P-atp2 promoter from the BioBrick Registry (BBa_K1675021) for the basic range. Each sequence was paired with a unique corresponding reporter sequence so that if each pH sensitive plasmid were in the same environment, the specific pH of the system could be seen. The reporters used were, BBa_E1010 for the CadC construct, BBa_K1033916 for the CpxA-CpxR complex, and BBa_K592009 for the P-atp2 promoter.
CadC
The CadC operon is a native pathway in E. coli, involved in the cadaverine synthesis pathway. The protein CadC protein on the operon is produced and activates segments downstream of the operon on the CadBA receptors. The CadC protein is pH sensitive to an external pH 5.5 and below, as well as lysine dependent. A point mutation on codon 265, in which argenine is converted to cystine, causes the CadC protein to become lysine independent (Dell, Neely, Olson, 1994).
Unfortunately, we have been unable to grow the modified CadC operon in E. coli suggesting some form of cell toxicity. Due to this apparent toxicity, no data regarding this mutant CadC could be collected. Alternative candidates are being explored for other pH sensors that sense in the acidic range.
CpxA-CpxR
CpxA-CpxR is a two-component mechanism that is activated at pH 7.4 and repressed at pH 6.0. CpxA is an intermembrane protein that autophosphorylates at a certain external pH, CpxR (a kinase) then gets phosphorylated by CpxA and acts as a transcription factor. This system originally is a transcription factor for the virF gene, but we replaced virF with the Reporter. The original sequence was found in Shigella sonnei, but E. coli has a homolog of these proteins so all that is required on the construct is the appropriate prefix/suffix and CpxR binding site.

The order from left to right is Control pH 6-9 and then Experimental pH 6-9. These are showing the gradient change in expression accordingly with the change of pH due to a pH-dependent promotor compared to consistent expression accordingly with a promoter that is always "on". The main point is that the Control at pH 6 has more expression of the Yellow-Green Chromoprotein than the Experimental at pH 6. The pH-Dependent promoter of the Experimental group is down-regulated at pH 6 whereas the control is not. Also, there is an increase in YGCP expression between the Experiment pH 7 and pH 8 that is not seen in the Control between pH 7 and pH 8. The normalized data is below shows the relative expression of YGCP. The construct can be found on the iGEM registry as: Bba_K2097000.

P-atp2
The P-atp2 promoter, native to the bacterium Corynebacterium glutamicum is reportedly induced at pH 7, to pH 9 (BIT-China-2015 and BBa_K1675021). Utilizing the blue chromoprotein (BBa_K592009), a test was designed in which a plasmid containing the P-atp2 promoter with the blue chromoprotein was grown alongside an E. coli line that contained a plasmid with just the blue chromoprotein. We expected to see constant blue chromoprotein production in the control series (those that lacked P-atp2) and a visual increase in blue chromoprotein as the pH was raised from 6 to 9 in the cells that contained the P-atp2 construct.

Next Steps and the GOX Sequences as Putative Promoters