Ianoverman (Talk | contribs) |
Ianoverman (Talk | contribs) |
||
| Line 24: | Line 24: | ||
</html> | </html> | ||
[[File:T--Austin_UTexas--pH_Dependent_Promoter.jpeg|thumb|left|600px| Insert caption here.]] | [[File:T--Austin_UTexas--pH_Dependent_Promoter.jpeg|thumb|left|600px| Insert caption here.]] | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
<html> | <html> | ||
Revision as of 05:38, 17 October 2016
Registry Parts
pH Sensors
CpxA-CpxR
CpxA-CpxR is a two-component mechanism that is activated at pH 7.4 and repressed at pH 6.0. CpxA is an intermembrane protein that autophosphorylates at a certain external pH, CpxR (a kinase) then gets phosphorylated by CpxA and acts as a transcription factor. This system originally is a transcription factor for the virF gene, but we replaced virF with the Reporter. The original sequence was found in Shigella sonnei, but E. coli has a homolog of these proteins so all that is required on the construct is the appropriate prefix/suffix and CpxR binding site.

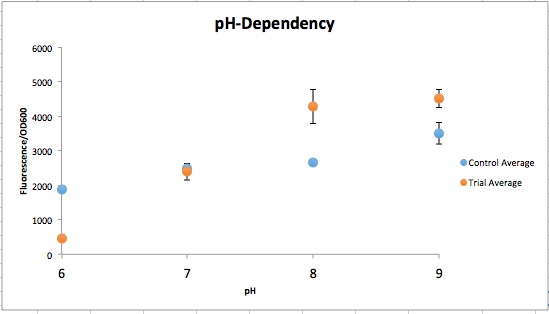
Figure 1 qualitatively depicts the Control at pH 6 with more expression of the Yellow-Green Chromoprotein than the Experimental at pH 6. The pH-Dependent promoter of the Experimental group is down-regulated at pH 6 whereas the control is not. Also, there is an increase in YGCP expression between the Experiment pH 7 and pH 8 that is not seen in the Control between pH 7 and pH 8. The normalized data in Figure 2 shows the relative expression of YGCP since the qualitative data is ambiguous. The construct can be found on the iGEM registry as: Bba_K2097000.
Next Steps and the GOX Sequences as Putative Promoters