Registry Parts
pH Sensors
CpxA-CpxR
CpxA-CpxR is a two-component mechanism that is activated at pH 7.4 and repressed at pH 6.0. CpxA is an intermembrane protein that autophosphorylates at a certain external pH, CpxR (a kinase) then gets phosphorylated by CpxA and acts as a transcription factor. This system originally is a transcription factor for the virF gene, but we replaced virF with the Reporter. The original sequence was found in Shigella sonnei, but E. coli has a homolog of these proteins so all that is required on the construct is the appropriate prefix/suffix and CpxR binding site.

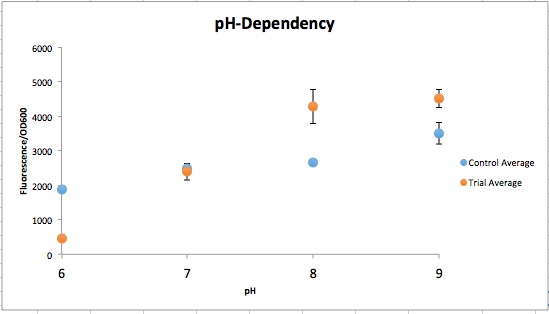
Figure 1 qualitatively depicts the Control at pH 6 with more expression of the Yellow-Green Chromoprotein than the Experimental at pH 6. The pH-Dependent promoter of the Experimental group is down-regulated at pH 6 whereas the control is not. Also, there is an increase in YGCP expression between the Experiment pH 7 and pH 8 that is not seen in the Control between pH 7 and pH 8. The normalized data in Figure 2 shows the relative expression of YGCP since the qualitative data is ambiguous. The construct can be found on the iGEM registry as: Bba_K2097000.
P-atp2
The P-atp2 promoter, native to the bacterium Corynebacterium glutamicum is reportedly induced at pH 7, to pH 9 (BIT-China-2015 and BBa_K1675021). Utilizing the blue chromoprotein (BBa_K592009), a test was designed in which a plasmid containing the P-atp2 promoter with the blue chromoprotein was grown alongside an E. coli line that contained a plasmid with just the blue chromoprotein. We expected to see constant blue chromoprotein production in the control series (those that lacked P-atp2) and a visual increase in blue chromoprotein as the pH was raised from 6 to 9 in the cells that contained the P-atp2 construct.
Next Steps and the GOX Sequences as Putative Promoters