| (4 intermediate revisions by the same user not shown) | |||
| Line 79: | Line 79: | ||
</br></br><h3 style="padding-top:0px;">Aim 1</h3> | </br></br><h3 style="padding-top:0px;">Aim 1</h3> | ||
<div class="col-md-14 content" style="max-width:1000px;padding:10px 50px;"> | <div class="col-md-14 content" style="max-width:1000px;padding:10px 50px;"> | ||
| + | We were able to design constructs for two therapeutic proteins, aprotinin and platelet derived growth factor (PDGF). Our goal was to use these therapeutic proteins to decrease wound healing times. Aprotinin, being a matrix-metalloprotease (MMP) inhibitor, will inhibit wound-site proteases, preventing these proteases from breaking down the healing extra-cellular matrix (ECM). PDGF will stimulate cellular regeneration in chronic wounds, helping to speed up and fortify the wound healing process. | ||
| + | </br></br> | ||
<center><img src="https://static.igem.org/mediawiki/2016/a/a4/T--Lubbock_TTU--dc2.png" width="350"></img> | <center><img src="https://static.igem.org/mediawiki/2016/a/a4/T--Lubbock_TTU--dc2.png" width="350"></img> | ||
| + |   | ||
<img src="https://static.igem.org/mediawiki/2016/0/09/T--Lubbock_TTU--dc4.png" width="350"></img></center> | <img src="https://static.igem.org/mediawiki/2016/0/09/T--Lubbock_TTU--dc4.png" width="350"></img></center> | ||
</br><div id="readmore"><a href="https://2016.igem.org/Team:Lubbock_TTU/Experiments">Experimental Data →</a></br></div> | </br><div id="readmore"><a href="https://2016.igem.org/Team:Lubbock_TTU/Experiments">Experimental Data →</a></br></div> | ||
| Line 92: | Line 95: | ||
</br></br><h3 style="padding-top:0px;">Aim 2</h3> | </br></br><h3 style="padding-top:0px;">Aim 2</h3> | ||
<div class="col-md-14 content" style="max-width:1000px;padding:10px 50px;"> | <div class="col-md-14 content" style="max-width:1000px;padding:10px 50px;"> | ||
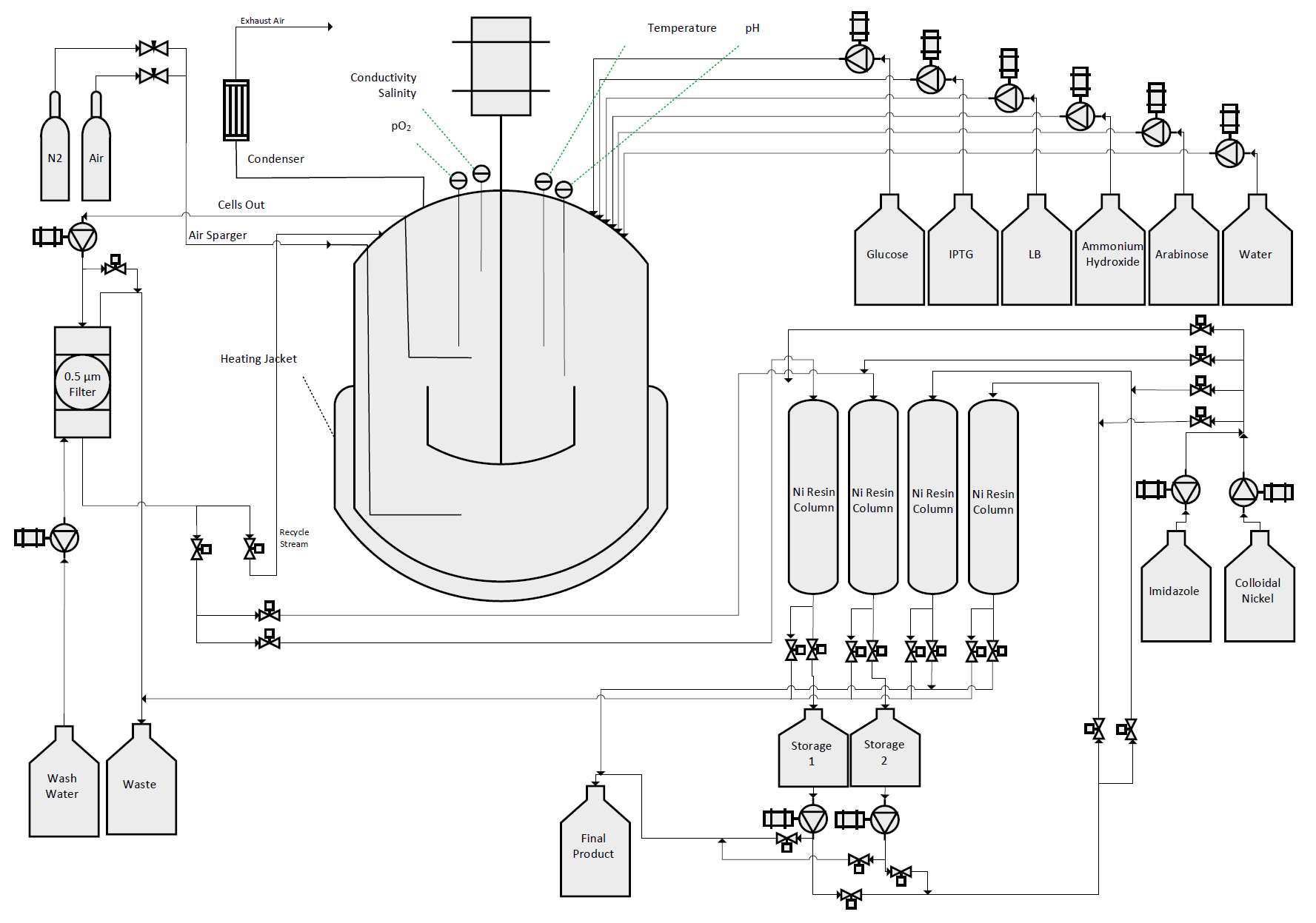
| + | We designed, created, and tested a bioreactor to grow our bacteria in conditions so as to maximize protein production. With this optimal environment, we hoped to create a sustainable, cost-effective protocol to mass-produce our therapeutic agents. | ||
| + | </br></br> | ||
<center><img src="https://static.igem.org/mediawiki/2016/e/e7/T--Lubbock_TTU--dc1.png" width="350"></img> | <center><img src="https://static.igem.org/mediawiki/2016/e/e7/T--Lubbock_TTU--dc1.png" width="350"></img> | ||
| + |   | ||
<img src="https://static.igem.org/mediawiki/2016/7/71/T--Lubbock_TTU--dc5.png" width="350"></img></center> | <img src="https://static.igem.org/mediawiki/2016/7/71/T--Lubbock_TTU--dc5.png" width="350"></img></center> | ||
</br><div id="readmore"><a href="https://2016.igem.org/Team:Lubbock_TTU/Experiments">Experimental Data →</a></br></div> | </br><div id="readmore"><a href="https://2016.igem.org/Team:Lubbock_TTU/Experiments">Experimental Data →</a></br></div> | ||
| Line 105: | Line 111: | ||
</br></br><h3 style="padding-top:0px;">Aim 3</h3> | </br></br><h3 style="padding-top:0px;">Aim 3</h3> | ||
<div class="col-md-14 content" style="max-width:1000px;padding:10px 50px;"> | <div class="col-md-14 content" style="max-width:1000px;padding:10px 50px;"> | ||
| + | We made a control collagen matrix to compare with a collagen matrix that will be infused with our purified proteins, aprotinin and PDGF, which would imitate the human extracellular matrix. This would give the wound bed all the resources it would need for a speedy recovery. | ||
| + | </br></br> | ||
<center><img src="https://static.igem.org/mediawiki/2016/8/85/T--Lubbock_TTU--dc3.png" width="350"></img></center> | <center><img src="https://static.igem.org/mediawiki/2016/8/85/T--Lubbock_TTU--dc3.png" width="350"></img></center> | ||
</br><div id="readmore"><a href="https://2016.igem.org/Team:Lubbock_TTU/Experiments">Experimental Data →</a></br></div> | </br><div id="readmore"><a href="https://2016.igem.org/Team:Lubbock_TTU/Experiments">Experimental Data →</a></br></div> | ||
| Line 117: | Line 125: | ||
</br></br><h3 style="padding-top:0px;">Conclusion</h3> | </br></br><h3 style="padding-top:0px;">Conclusion</h3> | ||
<div class="col-md-14 content" style="max-width:1000px;padding:10px 50px;"> | <div class="col-md-14 content" style="max-width:1000px;padding:10px 50px;"> | ||
| − | + | Coupling a bioreactor with our Kil protein secretion method, we will be able to purify large quantities of DsbA tagged therapeutic protein from the supernatant with a continuous flow through process. Western blot analysis helped us to confirm the identity of our purified PDGF-B protein, and the bioreactor allowed us to produce large amounts of our secretion proteins. | |
</div> <!-- End of col-md-14 --> | </div> <!-- End of col-md-14 --> | ||
</br></br><div class="col-md-2"></div> | </br></br><div class="col-md-2"></div> | ||
| Line 128: | Line 136: | ||
</br></br><h3 style="padding-top:0px;">Gold Medal Part Improvement</h3> | </br></br><h3 style="padding-top:0px;">Gold Medal Part Improvement</h3> | ||
<div class="col-md-14 content" style="max-width:1000px;padding:10px 50px;"> | <div class="col-md-14 content" style="max-width:1000px;padding:10px 50px;"> | ||
| − | + | A big part of the 2016 TTU iGEM teams project relied upon being able to produce Aprotinin (BBa_K1456006), a protease inhibitor for infusion into a collagen scaffold. To increase our chances we decided to improve upon this part which was created in 2014 by the ATOMS-Turkiye iGEM team. | |
| + | |||
| + | </br></br> | ||
| + | The first thing we did to improve this part was to remove two illegal sites to make it compatible with all iGEM RFC standards. Secondly, we found out that Aprotinin contains three disulphide bonds, which make correct folding in the cytoplasm difficult. To overcome this problem we decided to add a periplasmic localization peptide to K1456006, which allows for cotranslational translocation into the periplasm, where disulphide bonds are more easily formed. | ||
| + | |||
| + | </br></br> | ||
| + | <center><img src="https://static.igem.org/mediawiki/2016/7/7f/T--Lubbock_TTU--ptimpr.png" width="450"></img></center> | ||
| + | |||
</div> <!-- End of col-md-14 --> | </div> <!-- End of col-md-14 --> | ||
</br></br><div class="col-md-2"></div> | </br></br><div class="col-md-2"></div> | ||
Latest revision as of 03:28, 20 October 2016
Project Overview
In 2010 it was estimated that 6.5 million people in the United States alone suffered from chronic wounds, accruing an annual cost of approximately $2.5 billion. Furthermore, experts predict that the burden of chronic wounds will increase rapidly in the near future due to increasing medical costs, an aging population, and the emergence of antibiotic resistant bacteria. Chronic wounds are characterized by their inability to progress through an orderly set of stages within a time period of about three months. Wound healing progresses through four successive stages known as hemostasis, inflammation, proliferation and remodeling.



Our Goals
We will use synthetic biology principles to help treat chronic wounds by targeting the overproduction of wound site protease.
- For Aim 1 we will genetically engineer E. coli to produce a protease inhibitor and platelet derived growth factor.
- For Aim 2 we will purify the protease inhibitor and platelet derived growth factor in a bioreactor.
- For Aim 3 we will design a collagen bandage that mimics the human extracellular matrix, and infuse it with purified protease inhibitor and platelet derived growth factor.
Aim 1
We were able to design constructs for two therapeutic proteins, aprotinin and platelet derived growth factor (PDGF). Our goal was to use these therapeutic proteins to decrease wound healing times. Aprotinin, being a matrix-metalloprotease (MMP) inhibitor, will inhibit wound-site proteases, preventing these proteases from breaking down the healing extra-cellular matrix (ECM). PDGF will stimulate cellular regeneration in chronic wounds, helping to speed up and fortify the wound healing process.




Aim 2
We designed, created, and tested a bioreactor to grow our bacteria in conditions so as to maximize protein production. With this optimal environment, we hoped to create a sustainable, cost-effective protocol to mass-produce our therapeutic agents.



Aim 3
We made a control collagen matrix to compare with a collagen matrix that will be infused with our purified proteins, aprotinin and PDGF, which would imitate the human extracellular matrix. This would give the wound bed all the resources it would need for a speedy recovery.


Conclusion
Coupling a bioreactor with our Kil protein secretion method, we will be able to purify large quantities of DsbA tagged therapeutic protein from the supernatant with a continuous flow through process. Western blot analysis helped us to confirm the identity of our purified PDGF-B protein, and the bioreactor allowed us to produce large amounts of our secretion proteins.


