NOTEBOOK.
I should probably put some Table of Contents up here, so one could easily skip from day to day... at any rate, here's the current info dump.
Heads-up: I know there aren't circles on the list. This is apparently harder to do than I had envisioned?
6/13/16
- (HC) Made LB-Amp plates
- 450ml of DI water
- 5.0g tryptone
- 2.5g yeast
- 5.0g NaCl
- 7.5g Agar
- This was then adjusted to a pH of 7.0 using NaOH. It was then autoclaved. 2.5mL of 10mg/mL of Amp for a final concentration of 50 μg/mL was achieved.
6/14/16
- (HC) made CSM -His plates
6/15/16
- (XX) Goal: Run a gel to check whether pSB416 GPD has PstI site removed (diagnostic gel)
- 1. pSB416 GPD cut wit AvaI
- 2. pSB416 GPD cut with PstI
- 3. p146GPD cut with AvaI
- 4. p416GPD cut with PstI
- Major steps:
- Digest:
6 sterile water, 1 CutSmart buffer, 2 DNA, 1 enzyme (AvaI or PstI-HF)
- Gel Order: L, 1, 3, 2, 4
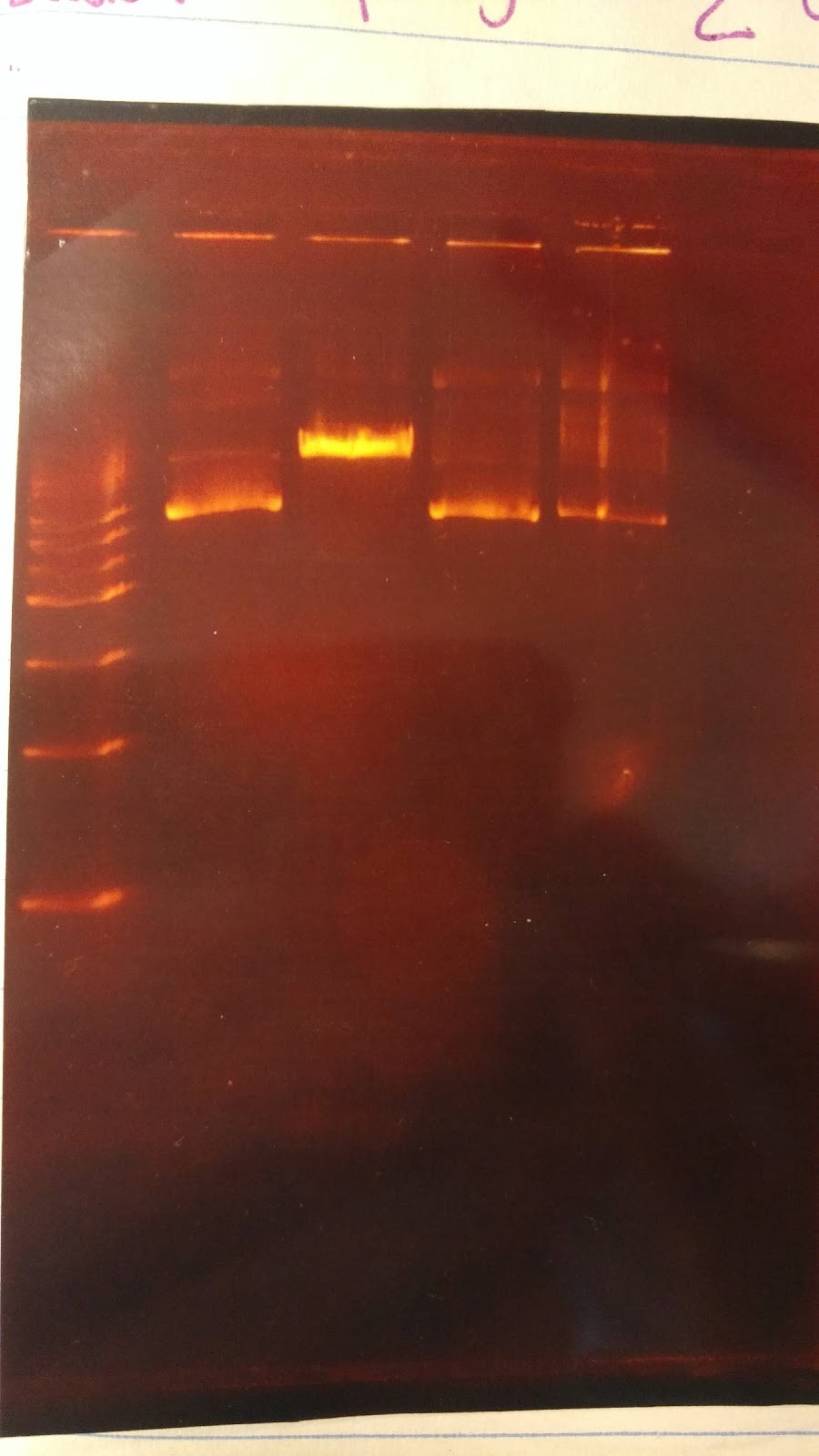
- Expected results on this diagnostic gel:
- 1. No cut because no AvaI site in pSB416 GPD
- 2. Should only see one cut because only one PstI site
- 3. Should see one cut
- 4. Should see two bands
- Experimental results:
Gel results are not good; need to rerun this diagnostic gel
- Other Activities:
- Created mRPS12+/KanMZ- (parent) and mRPS12-/KanMX+ (mutant) master plate on YPD
- Replica plating YPG (Should this be YPD???) , YPG, -His, -Ura, -Ura+Glycerol, YPD+Paro, YPG+Paro
6/16/16
(HC) Ran a boiling and spin prep on P413 GPD
- Boiling Prep Procedure:
- 1) Pellet cells from 1.5mL of an overnight L-Amp culture of transformed E. coli cells in a microcentrifuge for 20-30 seconds and remove supernatant.
- 2) Use a toothpick and vortexer to resuspend the cells in 100 μl(0.1mL) of STET buffer(8% sucrose; 5%Triton X-100;50 mM EDTA; Tris-Cl, pH 8.0) containing 1mg/mL lysozyme(prepare 1 mg/mLenzyme in STET just prior to use; 1mL= 10 preps).
- 3)Place the suspension in a boiling water bath for 90 seconds.
- 4) Spin in a microcentrifuge for 15 min at the highest speed.
- 5)Remove the pellet (a gelatinous mass of cell debris) with a toothpick and discard.
- 6) Precipitate nucleic acid at -20℃ for 30 minutes using an equal volume of isopropanol.
- 7) Spin the tube in a microcentrifuge for 10 minutes at RTo to pellet the nucleic acids.
- 8) Was pellet twice with 70% ethanol and dry thoroughly.
- 9) Resuspend the pellet in 50 μL of sterile DI water or TE buffer.
- Spin Prep: We used the QIAprep Spin Miniprep Kit to run a spin prep and followed the instructions inside.
- (XX)Goal: Rerun the diagnostic gel stated yesterday (6/15/16)
- Experimental results:
Good gel results, confirming that the PstI site has been removed in pSB416 GPD. pSB416 ready to be sequenced.
- Other activities:
Incubated pSBIC3 mRPS12 TU and pSBIC3 mRPS12 mls (from last year iGEM team??) in LB-chloramphenicol broth @37C overnight at 250 rpm
6/17/16
(HC) Ran two digests for P413 GPD
- Analytical Digest:
- 5 μL of DNA(Boiling Prep)
- 1μL of PstI HF
- 1μL of CutSmart buffer
- 3μL of sterile DI water
- Preparative Digest:
- 7μL of DNA(Spin Prep)
- 1μL of NheI
- 1μL of NsiI
- 1μL of CutSmart buffer
- Gel Results:
- PREP GEL
- Lane 1: Ladder
- Lane 3: Preparative Digest
- INSERT NOTEBOOK PICTURE #001 HERE
- Lane 1: Ladder
- Lane 2: Uncut (2μL of DNA and 8μL of water)
- Lane 3: Analytical Digest
- INSERT NOTEBOOK PICTURE #002 HERE
- (XX) Master plate results:
- Nothing grew on control plates (-His, -Ura, -Ura+Glycerol)
- Both parent and mutant grew on YPD and YPD+Paro, indicating that Paro has no effect on yeast growth
- Mutant cannot grow on YPG or YPG+Paro because of the mRPS12 gene knockout
- Other Activities:
- Cut p413GPD with NheI and NsiI to get rid of the illegal PstI site
- Massive gel to run boiling preps
- Conclusion:
- P413GPD cut looks like an uncut plasmid
- Massive gel has many blurry bands, which indicates that boiling preps are very impure
6/20/16
- (XX) Unltimate Goal:
- Put pSBIC3 mRPS12 TU into our own standardized yeast vectors
- Validate mls function by creating mls-yeGFP construct
- Goal for today:
- Digest and Harvest mRPS12 TU from pSBIC3(tube 1)
- 5μL of DNA
- 1μL of EcoRI-HF
- 1μL of pstI-HF
- 1μL of CutSmart buffer
- 2μL of sterile water
- Digest and Harvest mRPS12 mls from pSBIC3 (tube 2)
- 5μL of DNA
- 1μL of EarI
- 1μL of SpeI
- 1μL of CutSmart buffer
- 2μL of sterile water
- Gel Order: L, 1C, 1XX, 1XL, 2C, 2XX, 2XL
- **C-uncut plasmid, XX-Xintong, XL-Xander
- Gel results:
- Uncut does not show up on the gel
- Not a very good results but we harvested the bands anyway
- Other activiites:
- Excise bands from the gel and perform gel extraction (QIA kit):
- Tube 1 = 0.26 g
- Tube 2 = 0.14 g
- Tube 3 = 0.17 g
- *tube 3 contains p413 GPD with NsII/NheI cut
- Prepare Yeast Competent Cells
- 10 mL YPD, inoculated at 250RPM @ 30C
- Both BY4741 and BY4741 mRPS12 is prepared
6/21/16
- (XX) Goal for Today:
- DNA Assembly
- Major Steps:
- We used nanodrop to determine the amount of DNA in tube 1 (TU) and tube 2 (mls)
- Tube 1: 4.5 ng/
- Tube 2: 4.0 ng/
- * Nano drop results indicate that we need to precipitate DNA for further experiment
- After precipitation, nanodrop reports 12.6 ng/ DNA
- NEBuilder HiFi DNA Assembly Cloning Kit:
- 2μL Ear/SpeI digest pSBIC3 mls
- 8μL fragment 16-4 (yeGFP)
- 10μL HiFi Master
- *All three above, set the reaction on ice; 1:2 vecotr: insert
- Thermocycler
- Other Activities:
- Yeast competent cell from yesterday did not grow
- Prepare new yeast competent cells: 5 mL YPD in round bottom tubes
6/22/16
- (XX)Goal:
- Prepare Yeast Competent cells
- Diagnostic gel on mRPS12 TU minipreps
- Major Steps:
- Yeast competent cells:
- Spectromphotometer OD 260:
- 1000 YPD Blank
- 1000 KnockOut(KO) has 1.777 Abs.
- 1000 Parent has 2.222 Abs.
- Regrow these cells into 15 mL broth for about 2 hrs:
- Take 0.5 mL current culture into 14.5 mL YPD
- Spectrophotometer Results:
- KO: 0.390 Abs.
- Parent: 0.692 Abs.
- Follow yeast competent cell protocol
- Diagnostic Gel:
- 1: pSBIC3 mRPS12 TU miniprep 1 cut with EcoRI and PstI
- 2: pSBIC3 mRPS12 TU miniprep 2 cut with EcoRI and PstI
- 3: pSBIC3 mRPS12 TU miniprep 1 cut with PstI
- 4: pSBIC3 mRPS12 TU miniprep 2 cut with PstI
- Gel Order: L, 1, 2, 3, 4
- Gel Results:
- For #2, fragment should be around 500bp
- Gel results indicate that we should redo minipreps or bad enzymes
6/23/16
- (XX) Goal:
- Test if the enzymes are still okay to work with
- Redo miniprep at the same time
- Major steps:
- Enzyme test:
- Tube 1: EcoRI-HF
- Tube 2: PstI-HF
- Tube 3: SpeI-HF
- Tube 4: EarI
- Gel Order: 1, 2, 3, 4, L
- Gel Results: enzymes are working properly
6/24/16
- (HC) After extracting P413 GPD fragment from the gel, I precipitated the DNA
- Precipitating DNA:
- 1.)Add 1-2 μL of 5M NaCl to 30 μL of DNA
- 2.)Add 62 μL of ice cold EtOH
- 3.)Let sit on ice for 30 minutes
- 4.)Centrifuge @ 0℃ for 10 minutes at 13 krpm
- 5.)Remove supernatent
- 6.)Fill tube to halfway point with 70% EtOH
- 7.)Centrifuge @ 4℃ for 2 minutes
- 8.)Remove supernant
- 9.)Let dry
- 10.)Add 6μL of Sterile Water.
- Obtained concentration of DNA with NanoDropper
| Nucleic Acid Concentratione | A @ 260 nm | A @ 280 nm | 260/280 | 260/230 |
|---|---|---|---|---|
| 75.7 ng/μL | 1.514 | 0.922 | 1.64 | .033 |
- (JM/CH)
-
Today we are starting the day off by running a gel.
We just called iGEM to confirm that we can indeed submit vectors as parts
The gel we are running today is to confirm that we only have 1 PstI site in all of our plasmids
The hope: to see one solid band from all of our plasmids except for the controls, the controls should be two bands because they still have two PstI sites
Another thing we are doing today is we are making media LB Chloro.
O

