|
|
| (19 intermediate revisions by 5 users not shown) |
| Line 1: |
Line 1: |
| | {{:Team:Tianjin/Templates/MaterialTheme|}} | | {{:Team:Tianjin/Templates/MaterialTheme|}} |
| | | | |
| − | {{:Team:Tianjin/Templates/AddCSS|:Team:Tianjin/Note/Consortium/camarts.css}} | + | {{:Team:Tianjin/Templates/AddCSS|:Team:Tianjin/Note/R-R/camarts.css}} |
| | {{:Team:Tianjin/Templates/AddCSS|:Team:Tianjin/Experiment/css/font.css}} | | {{:Team:Tianjin/Templates/AddCSS|:Team:Tianjin/Experiment/css/font.css}} |
| − |
| |
| | <html lang="en"> | | <html lang="en"> |
| | + | <style> |
| | + | #Notes{ |
| | + | color: inherit; |
| | + | background-color: rgba(255, 255, 255, 0.1); |
| | + | } |
| | + | </style> |
| | + | |
| | <head> | | <head> |
| | <meta charset="utf-8" /> | | <meta charset="utf-8" /> |
| Line 12: |
Line 18: |
| | | | |
| | <meta content="width=device-width, initial-scale=1.0, maximum-scale=1.0, user-scalable=0" name="viewport" /> | | <meta content="width=device-width, initial-scale=1.0, maximum-scale=1.0, user-scalable=0" name="viewport" /> |
| | + | |
| | + | |
| | + | |
| | + | <style> |
| | + | #Notes{ |
| | + | color: inherit; |
| | + | background-color: rgba(255, 255, 255, 0.1); |
| | + | } |
| | + | </style> |
| | | | |
| | <!------------------------------------------- This part should be dismissed in WIKI u | | <!------------------------------------------- This part should be dismissed in WIKI u |
| Line 68: |
Line 83: |
| | <article id="post-4252" class="post-4252 post type-post status-publish format-standard has-post-thumbnail hentry category-150 tag-174 tag-xinjiang tag-173"> | | <article id="post-4252" class="post-4252 post type-post status-publish format-standard has-post-thumbnail hentry category-150 tag-174 tag-xinjiang tag-173"> |
| | <header class="entry-header"> | | <header class="entry-header"> |
| − | <div class="entry-title" align="center" >Notes For Modified Cyanobacteria:A Controllable Lipid Producer</div> | + | <div class="entry-title" align="center" >Notes For Cyanobacteria</div> |
| | </header><!-- .entry-header --> | | </header><!-- .entry-header --> |
| | | | |
| Line 133: |
Line 148: |
| | | | |
| | | | |
| − | <div align="center">
| |
| − | <figure>
| |
| − | <a href="https://static.igem.org/mediawiki/2016/3/37/Igem--6803-week2.jpg" data-lightbox="no" data-title="Fig.3 Proposed degradation pathway for TPA<sup>[2]</sup>"><img src="https://static.igem.org/mediawiki/2016/3/37/Igem--6803-week2.jpg" width="100%"></a>
| |
| − | <figcation>Fig.3 Proposed degradation pathway for TPA<sup>[2]</sup></figcaption>
| |
| − | </figure>
| |
| − | </div>
| |
| | | | |
| | | | |
| Line 145: |
Line 154: |
| | | | |
| | | | |
| − | <a href="https://static.igem.org/mediawiki/2016/3/37/Igem--6803-week2.jpg" data-lightbox="no"> | + | <a href="https://static.igem.org/mediawiki/2016/3/37/Igem--6803-week2.jpg" data-lightbox="no" data-title="7942"> |
| | <img align="center" src="https://static.igem.org/mediawiki/2016/3/37/Igem--6803-week2.jpg" alt="Igem-6803-week2" width="300px"><br/><br/> | | <img align="center" src="https://static.igem.org/mediawiki/2016/3/37/Igem--6803-week2.jpg" alt="Igem-6803-week2" width="300px"><br/><br/> |
| | </a> | | </a> |
| Line 194: |
Line 203: |
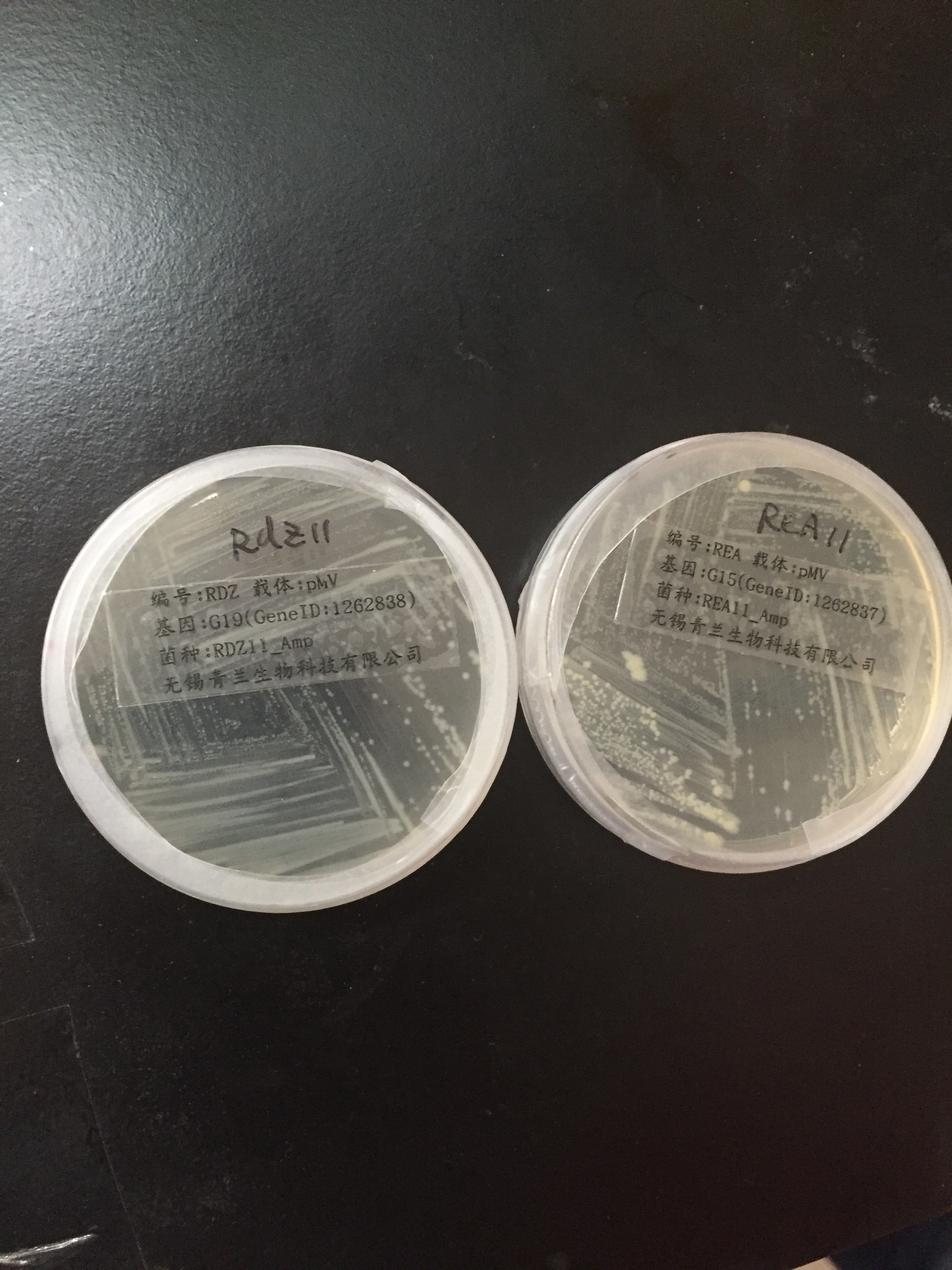
| | <div style="padding-left:32px;"><li>pMV-G19 and pMV-G15 containing our target genes were received.</li><br/> | | <div style="padding-left:32px;"><li>pMV-G19 and pMV-G15 containing our target genes were received.</li><br/> |
| | | | |
| | + | <a href="https://static.igem.org/mediawiki/2016/6/6d/Igem-6803-week3-1.jpg" data-lightbox="no" data-title="pMV-G19 and pMV-G15"> |
| | <img align="center" src="https://static.igem.org/mediawiki/2016/6/6d/Igem-6803-week3-1.jpg" alt="Igem-6803-week3-1" width="300px" ><br/><br/> | | <img align="center" src="https://static.igem.org/mediawiki/2016/6/6d/Igem-6803-week3-1.jpg" alt="Igem-6803-week3-1" width="300px" ><br/><br/> |
| | </a> | | </a> |
| Line 235: |
Line 245: |
| | <li>The enzyme-digested product was dephosphorylation.</li> | | <li>The enzyme-digested product was dephosphorylation.</li> |
| | <br/> | | <br/> |
| | + | |
| | + | <a href="https://static.igem.org/mediawiki/2016/a/aa/Igem-6803-week3.png" data-lightbox="no" data-title=" "> |
| | <img align="center" src="https://static.igem.org/mediawiki/2016/a/aa/Igem-6803-week3.png" alt="Igem-6803-week3-2" width="800" height="533"><br/> | | <img align="center" src="https://static.igem.org/mediawiki/2016/a/aa/Igem-6803-week3.png" alt="Igem-6803-week3-2" width="800" height="533"><br/> |
| | </a> | | </a> |
| Line 325: |
Line 337: |
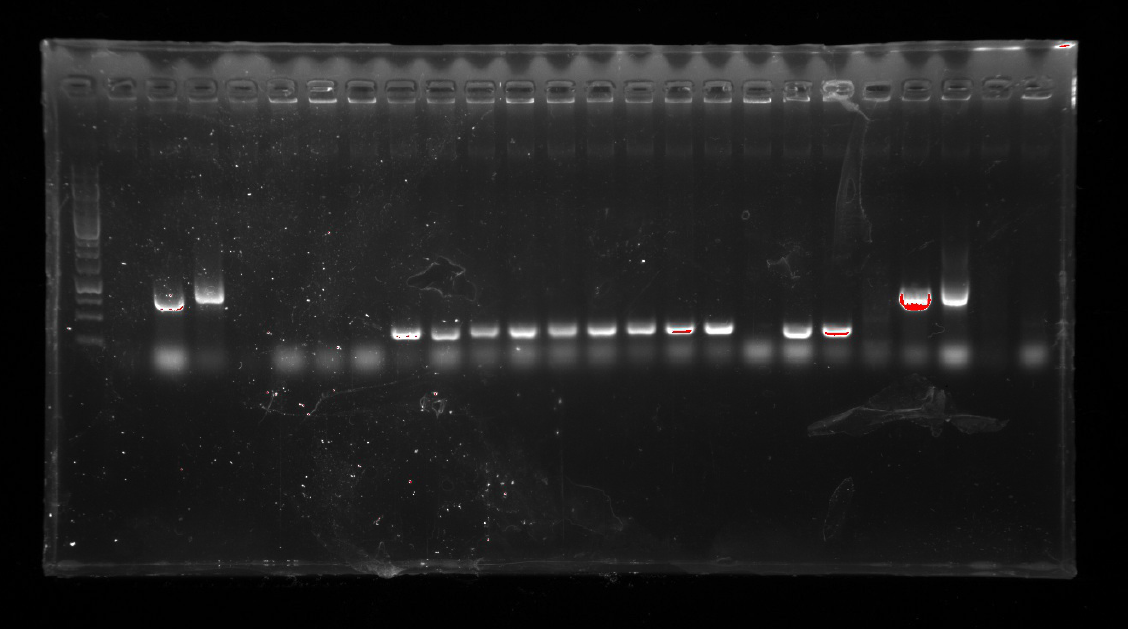
| | Two of the successful ones were used to inoculate overnight cultures.<br/> | | Two of the successful ones were used to inoculate overnight cultures.<br/> |
| | <br/> | | <br/> |
| | + | |
| | + | <a href="https://static.igem.org/mediawiki/2016/8/81/Igem-6803-week4.png" data-lightbox="no" data-title=" "> |
| | <img align="center" src="https://static.igem.org/mediawiki/2016/8/81/Igem-6803-week4.png" alt="Igem-6803-week4" width="800" height="533"> <br/> | | <img align="center" src="https://static.igem.org/mediawiki/2016/8/81/Igem-6803-week4.png" alt="Igem-6803-week4" width="800" height="533"> <br/> |
| | </a> | | </a> |
| Line 398: |
Line 412: |
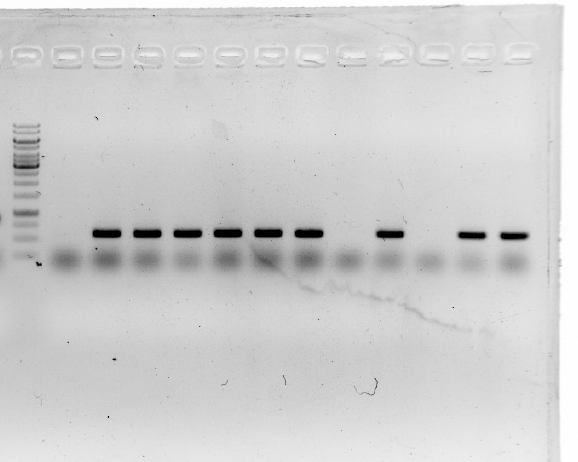
| | <b>The fourth and sixth ones were not successful.</b><br/> | | <b>The fourth and sixth ones were not successful.</b><br/> |
| | <br/> | | <br/> |
| | + | |
| | + | <a href="https://static.igem.org/mediawiki/2016/4/40/Igem-6803-week5.png" data-lightbox="no" data-title=" "> |
| | <img align="center"src="https://static.igem.org/mediawiki/2016/4/40/Igem-6803-week5.png" alt="Igem-6803-week5" width="800" height="533" ><br/> | | <img align="center"src="https://static.igem.org/mediawiki/2016/4/40/Igem-6803-week5.png" alt="Igem-6803-week5" width="800" height="533" ><br/> |
| | </a> | | </a> |
| Line 454: |
Line 470: |
| | | | |
| | <b><h1 style="font-size:108%"> Sep.22th</b></h1> | | <b><h1 style="font-size:108%"> Sep.22th</b></h1> |
| − | <div style="padding-left:32px;"><li>Gel electrophoresis showed that amplification of fragments was successfull.</li> | + | <div style="padding-left:32px;"> |
| | + | <li>Gel electrophoresis showed that amplification of fragments was successfull.</li> |
| | <li>Ligated <i>13</i> and <i>19-15</i> via overlap PCR.</li> | | <li>Ligated <i>13</i> and <i>19-15</i> via overlap PCR.</li> |
| | <li>This PCR worked well and the products were extracted from the gel using a Agarose Gel DNA Extration Kit.</li> | | <li>This PCR worked well and the products were extracted from the gel using a Agarose Gel DNA Extration Kit.</li> |
| | <li>Restriction digest on pCPC-3031-Ni with Sac I.</li> | | <li>Restriction digest on pCPC-3031-Ni with Sac I.</li> |
| | <br/> | | <br/> |
| − | <img align="center" src="https://static.igem.org/mediawiki/2016/8/89/Igem-6803-week6-2.jpg" alt="Igem-6803-week6-1" width="300px"><br/> | + | <a href="https://static.igem.org/mediawiki/2016/8/89/Igem-6803-week6-2.jpg" data-lightbox="no" data-title=" "> |
| − | <br/>
| + | <img align="center" src="https://static.igem.org/mediawiki/2016/8/89/Igem-6803-week6-2.jpg" alt="Igem-6803-week6-1" width="300px"> |
| | + | </a> |
| | + | |
| | <li>The fragment <i>13-19-15</i> was ligated onto T vector and transformed into E.coli via heat shock.</li> | | <li>The fragment <i>13-19-15</i> was ligated onto T vector and transformed into E.coli via heat shock.</li> |
| − | <br/> | + | |
| − | <img align="center" src="https://static.igem.org/mediawiki/2016/0/06/Igem-6803-week6.jpg" alt="Igem-6803-week6-2" width="300px" ><br/> | + | <a href="https://static.igem.org/mediawiki/2016/0/06/Igem-6803-week6.jpg" data-lightbox="no" data-title=" "> |
| | + | <img align="center" src="https://static.igem.org/mediawiki/2016/0/06/Igem-6803-week6.jpg" alt="Igem-6803-week6-2" width="300px" > |
| | + | </a> |
| | <br/> | | <br/> |
| | </div> | | </div> |
| Line 516: |
Line 537: |
| | <li>Three colonies were proved to be true. And two of them were used to inoculate overnight cultures.</li> | | <li>Three colonies were proved to be true. And two of them were used to inoculate overnight cultures.</li> |
| | <br/> | | <br/> |
| | + | <a href="https://static.igem.org/mediawiki/2016/2/2d/Igem-6803-week6-3.png" data-lightbox="no" data-title=" "> |
| | <img align="center" src="https://static.igem.org/mediawiki/2016/2/2d/Igem-6803-week6-3.png" alt="Igem-6803-week6-3" width="800" height="533" ><br/> | | <img align="center" src="https://static.igem.org/mediawiki/2016/2/2d/Igem-6803-week6-3.png" alt="Igem-6803-week6-3" width="800" height="533" ><br/> |
| | + | </a> |
| | <br/> | | <br/> |
| | | | |
| | </div> | | </div> |
| | <br/> | | <br/> |
| | + | |
| | + | |
| | | | |
| | <b><h1 style="font-size:108%"> Sep.26th</b></h1> | | <b><h1 style="font-size:108%"> Sep.26th</b></h1> |
| Line 527: |
Line 552: |
| | <br/> | | <br/> |
| | </div> | | </div> |
| | + | </br> |
| | + | |
| | + | |
| | + | |
| | + | <b><h1 style="font-size:108%"> Sep.28th</b></h1> |
| | + | <div style="padding-left:32px;"><li>We transformed the expression vector, pCPC-3031-Ni-13-19-15, into <i>Synechocystis 6803 </i>via electroporation.</li> |
| | <br/> | | <br/> |
| | + | <b>The story of 6803 is not over. Other team members will show the completed process during the presentation. </br> |
| | + | I will cherish the time spending with you for my whole life. ———by Jiawei Chen and Liangyu Qian</b> |
| | + | </div> |
| | | | |
| | | | |
| Line 556: |
Line 590: |
| | | | |
| | </div> | | </div> |
| − | <div class="side-nav">
| + | |
| − | <ul>
| + | |
| − | <li><a class="topLink" href="#Topp" style="color:#5555FF">Top</a></li>
| + | |
| − | <li><a class="topLink" href="#Week1" style="color:#5555FF">Week1</a></li>
| + | |
| − | <li><a class="topLink" href="#Week2" style="color:#5555FF">Week2</a></li>
| + | |
| − | <li><a class="topLink" href="#Week3" style="color:#5555FF">Week3</a></li>
| + | |
| − | <li><a class="topLink" href="#Week4" style="color:#5555FF">Week4</a></li>
| + | |
| − | <li><a class="topLink" href="#Week5" style="color:#5555FF">Week5</a></li>
| + | |
| − | <li><a class="topLink" href="#Week6" style="color:#5555FF">Week6</a></li>
| + | |
| − | <li><a class="topLink" href="#Week7" style="color:#5555FF">Week7</a></li>
| + | |
| − |
| + | |
| − | </ul>
| + | |
| − | </div>
| + | |
| | | | |
| | | | |