Protein Engineering
Motivation
The degradation of PET is completed in two steps by PETase and MHETase. PETase hydrolyze PET to mono(2-hydroxyethyl) terephthalic acid (MHET), which will be further decomposed by MHETase into two monomers, terephthalic acid (TPA) and ethylene glycol (EG)[1] . In this two-step reaction system, MHET, the product of PETase-mediated hydrolysis of PET, was found to be a very minor component, which reveals rapid MHET metabolism[1], indicating the rate determining step in this reaction is the first step, hydrolysis of PET. So in order to accelerate the whole PET degradation speed, increasing the activity of PETase is rather crucial. To enhance PETase hydrolysis activity, we first tried to understand the mechanism of the hydrolysis reaction by generally confirming active sites of PETase and 3 dementional structure simulation.
Serine-based Catalytic Triad Mechanism & 3D Model Simulation
Since there is no x-ray structure for PETase , the mechanism of PETase hydrolysis activity can not be exactly identified. But the binding site and catalytic site can be generally inferred according to α/β hydrolase mechanism. Based on the efforts have been made to identify and characterize bacterial cutinases[2], α/β hydrolase fold family contain a highly conserved characteristic GXSXG motif. With sequence analysis, PETase was also found to contain an accordant GWSMG motif. Meanwhile, we simulated a best fit model for PETase by SWISSMODEL, an automated comparative protein modeling server[3]. The template was Thc_Cut2, which shares 52% sequence identity with PETase[4]. As expected, the homology model of PETase displays a canonical α/β hydrolase fold with a Ser161-His237-Asp237 catalytic triad and a preformed oxyanion hole (Fig.1), suggesting a classic serine hydrolase mechanism.

Fig.1 Simulated 3D structure for PETase
Ribbon diagram of a predicted PETase model. The catalytic triad residues are shown as ball-and-sticks in green, formed by Ser161, His237 and Asp237, and the oxyanion hole binding site residues are in blue, formed by the main chain amides of Met161 and Tyr87.
Mutation Design Rationales
1.Hydrophobicity & space factor
Given that the hydrolysis of PET is a heterogeneous reaction, a prerequisite for its efficient reaction is an adequate interaction of the enzyme with the substrate at the solid-liquid interface. Since the substrate PET is hydrophobic polymer, both the hydrophobicity of the enzyme surface and the space around active site to accommodate PET may be the main factors for more efficient protein attachment and reaction activity[6]. In fact, tailoring enzymes for a more hydrophobic and larger substrate active site has been shown to facilitate PET hydrolysis by a fungal polyester hydrolase[5]. Also Silva et al. obtained a double mutant (Q132A/T101A) exhibiting a 2-fold increased hydrolysis activity against PET fibers by the substitution of spacious residues with alanine in its substrate binding site[6]. So we aimed to take the same strategy to modify hydrophobicity of the surface of PETase as well as broaden space near the active site to enhance the interaction between PETase and substrate.
2.Electrostatics factor
The electrostatic surface property in the vicinity to the active site was also found to be responsible for differences in hydrolysis efficiency. Exchange of the positively charged amino acids to the non-charged ones strongly increase the hydrolysis activity. In contrast, exchange of the uncharged amino acids by the negatively charged ones can lead to a complete loss of hydrolysis activity on PET films[7]. Changing the surface property is another strategy we took.
3.Conserved Residues
In another study, Wei and coworkers exchanged selected amino acid residues of TfCut2 involved in substrate binding with those in LC-cutinase(LCC) to relieve product inhibition and obtained enzyme variants with increased PET activity at 65 [8]. In the inspiration of their work as well as that the position of catalytic triad hold constant in different mature PET hydrolases with signal peptide excluded, we exchanged amino acid residues on the surface with highly conserved residues of the same position in other PET hydrolases, which can be a way to either increase the enzyme efficiency or confirm the essential sites which made PETase more efficient than other PET hydrolase.

Fig.2 Hydrophobicity of PETase surface The deeper the bule is, the more hydrophobic the protein surface is; On contrast, the deeper the brown is, the more hydrophilic the protein surface is.
Based on the above rationales, our general strategies of mutation design are as follows:
1).According to the 3D structure of PETase we simulated, which shows the hydrophobicity of protein surface (Fig.2), we changed the hydrophilic amino acid residues on the surface with hydrophobic ones to enhance the surface hydrophobicity;
2). Modify the electrostatics of its surface by changing charged amino acid residues to the uncharged ones;
3). Based on 3D structure predicted and docking simulation with substrate, we found several spacious residues adjacent to active site and binding site, which may inhibit the interaction of protein and substrate due to its large body. We substituted them with smaller amino acids to expose the catalytic site and binding site;
4). We generated a multiple sequence alignment by ClustalX with seven serine hydrolases from NCBI conserved with GXSXG motif and catalytic triad and reported to be able to degrade PET (Table.1). Based on the multiple sequence alignment result(Fig.3), we exchanged some essential residues adjacent to active site and binding site with the conserved residues of the same position in the multiple sequences.

Table.1 Enzymes in multiple sequence alignmentr
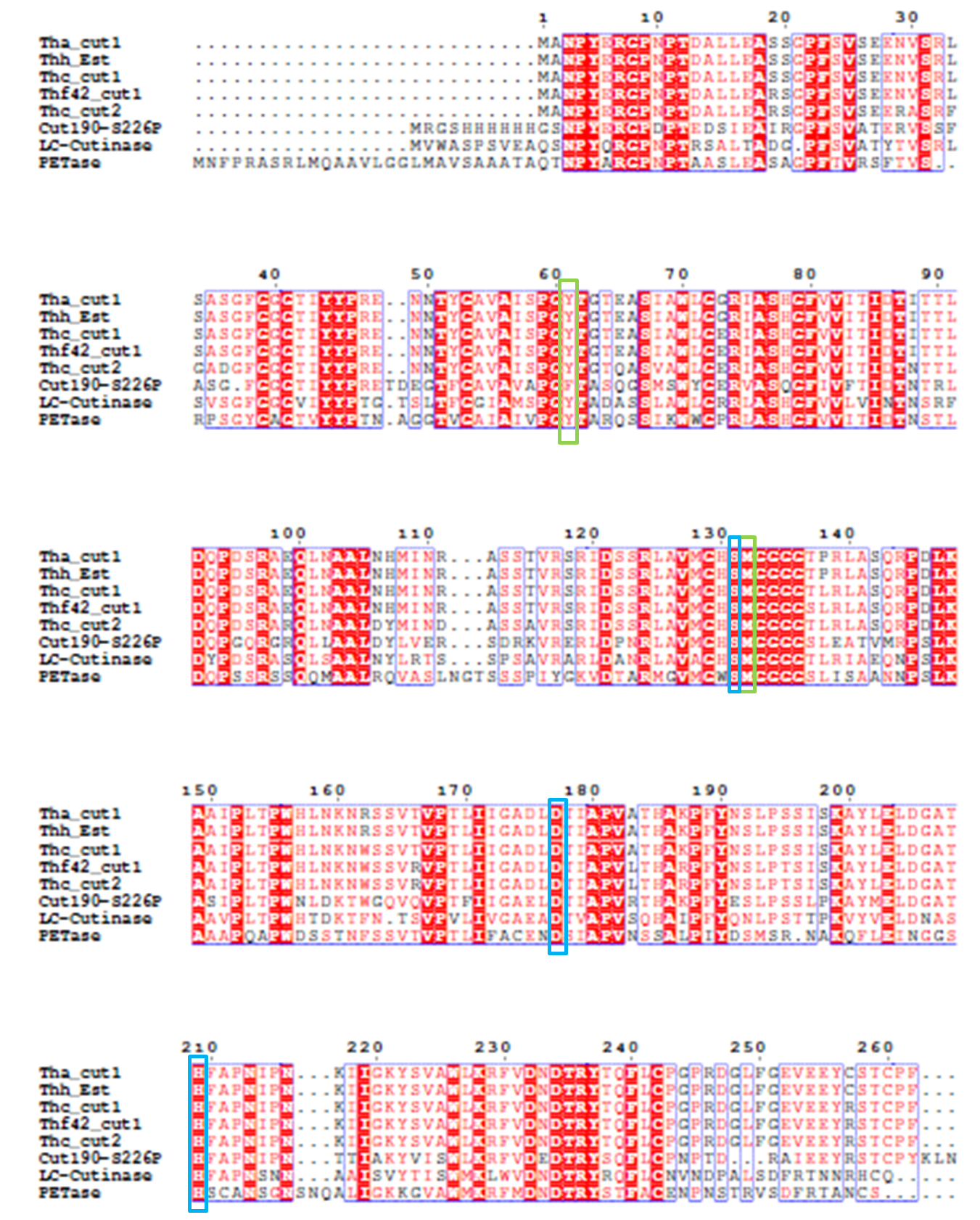
Fig.3 Multiple Sequence Alignment
The catalytic triad is shaded in blue box and the binding site is shaded in green box. The position of catalytic triad hold constant throughout the multiple sequences in mature protein with signal peptide excluded.
Mutants
Based on the Rationales above, we designed 22 potential mutations (Table.2) in purposes to enhance the efficiency of PETase hydrolysis activity and confirm potential determining site of PETase hydrolysis.
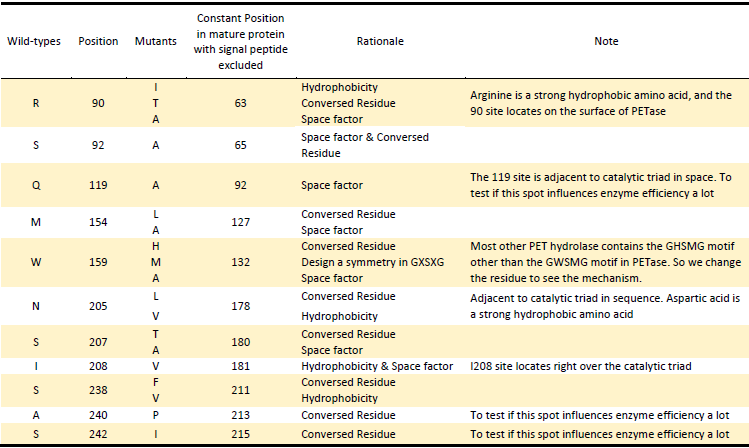
Table.2 Mutants of PETase
Site-directed mutagenesis
Site-directed mutagenesis was performed using recombinant PCR technique[9]. Our modified approach is based on the PCR amplification of target PETase fragment by mutagenic primers divergently oriented but overlapping at their 5’ ends. The mutagenic nucleotides are located in both forward and reverse primers. The approach contained two rounds of PCR. In the PCR Round I, we use PETase fragment as template, respectively use original forward primer of PETase & mutagenic reverse primer, and original reverse primer & mutagenic forward primer as primers. The product of PCR Round I are portions of PETase gene with an overlapping region. In the PCR Round II, we use products of PCR Round I as two templates and original primers as primers to generate the final mutant fragment (Fig.4). And each specific mutation was verified by sequencing.

Fig.4 Site-directed Mutagenesis
Assay system--CFPS
1.Overview
The CFPS system is the assay system in our project for the enzyme modification. The ability to produce a functional protein in the test tube, rather than in cells, is the essence of cell-free protein synthesis (CFPS)[11]. .

Fig.5. The basic process of CFPS system[2]
The lack of high-throughput approaches for expression and screening of large enzyme libraries is a major bottleneck for current enzyme engineering efforts. To address this need, some researchers[12] have developed a high-throughput, fluorescence-based approach for rapid one-pot, microscale expression, and screening of different kinds of enzymes. To go further, we try to make our effert to achieve integration of cell-free protein expression with activity screening of enzymes( site-directed mutants of PETase). .

Fig.6.One-pot approach for integrated expression and activity screening of enzymes
2.Background
At its core, synthetic biology is inspired by the power and diversity of the living world. It is an endeavor predicated on the idea that we can learn to more reliably and rapidly engineer biological function for compelling applications in medicine, biotechnology, and green chemistry. What is unique to synthetic biology is the application of an engineering-driven approach to accelerate the design-build-test loops required for reprogramming existing, and constructing new, biological systems. This is needed because the current approach to engineering cells is often extremely laborious, costly, and difficult. So far, efforts in synthetic biology have largely focused on the construction and implementation of genetic circuits, biological modules (compilations of biological ‘‘parts’’ that perform defined functions, e.g., ribosome binding sites, promoters, and genes), and synthetic pathways into genetically reprogrammed organisms. However, a major challenge exists because of our incomplete knowledge of how life works, the daunting complexity of cells, the unintended interference between native and synthetic parts, and (unlike typical engineered systems) the fact that cells evolve, have noise, and have their own agenda such as growth and adaptation. The guiding question, therefore, is how do we develop a new way of engineering in the face of these unique and complex features of biology? In other words, what is the suite of foundational technologies required for taming the complexity of living systems? One foundational technology that has emerged is the ability to construct new programs that carry out user-defined functions from ‘‘parts’’ of DNA (e.g., biological switches or oscillators). Biological computer-aided design (BioCAD) tools, directed evolution, and cell-free systems will also each have a role. Although in vivo synthetic biology projects are dominant in the field, we focus this review on the role of cell-free synthetic biology. Without the need to support ancillary processes required for cell viability and growth, cell-free systems offer a powerful platform for accelerating the optimization of synthetic pathways and for not only harnessing but also expanding the chemistry of life.
The power of cell-free systems was first appreciated over a hundred years ago. In 1897, Eduard Buchner used yeast extract to convert sugar to ethanol and carbon dioxide, for which he won the Nobel Prize (1907 Chemistry). Since then, cell-free systems have continued to play an important role in our understanding of fundamental biology. For example, Nirenberg’s discovery of the genetic code, for which he received the Nobel Prize in 1968, shed light into how protein synthesis works. Cell-free systems have since continued to unravel other fundamental discoveries such as the understanding of eukaryotic translation. Beyond using cell-free systems for biochemical analysis, the successful recapitulation of biological function in vitro has inspired attempts to use cell-free systems for product synthesis. Indeed, there are more than 30 years of successful industrial history with biotransformations in vitro. For the most part, in vitro biocatalytic transformation has focused on single transformations, such as enabling the production of specific chirality. In another example, single-enzyme cell-free systems, such as cell-free DNA replication (the polymerase chain reaction (PCR)) are ubiquitous to molecular biology labs and have revolutionized modern biological research. PCR in combination with other common molecular tools such as DNA restriction digest and ligation reactions allow us to assemble DNA sequences at will. In other approaches, crude extract cell-free systems (CECFs) and synthetic enzymatic pathways (SEPs) have practical applications in protein and peptide synthesis and evolution, small molecule production, cellular/metabolic pathway investigation, and non-natural product synthesis.
As one of the most common examples of CECFs, cell-free protein synthesis (CFPS) provides a useful starting point for examining the utility of cell-free systems. To produce target proteins of interest, CFPS systems utilize ensembles of biocatalysts that carry out translation from crude extracts derived from bacterial, plant, or animal cells. Upon incubation with essential amino acids and energy substrates, the ensemble of activated catalysts within the cell lysate acts as a chemical factory to polymerize amino acids into polypeptides. Although any organism can be used to provide a source of crude lysate, the most productive CECFs are derived from Escherichia coli extract[13].
The translation machinery is the engine of life. In E.coli CFPS the translation machinery is typically about 20-fold more dilute than in thecell, decreasing the rates of initiation, elongation and protein accumulation . As well, the average distance between two adjacent ribosomes on a single mRNA strand increases and polysomes are less likely to form. Despite these differences, CFPS can benefit from the relative slower synthesis rate and the distance between ribosomes by allowing nascent polypeptide chain more time and space to form desirable intra-peptide chain contacts, while decreasing the probability for undesirable, non-specific inter-peptide chain contacts, thereby increasing the probability of proper folding and decreasing the probability of aggregation.
Cell-free protein synthesis is a widely used method in molecular biology. Production of proteins using cell-free protein synthesis usually takes a few hours, in contrast to production of proteins in cells, which typically takes days to weeks. In fact, even first-time users can often obtain newly synthesized proteins in one day using a commercial system.
The diversity of the cell-free systems allows in vitro synthesis of a wide range of proteins for a variety of downstream applications, such as screeening of enzymes activities. In the post-genomic era, cell-free protein synthesis has rapidly become the preferred approach for high-throughput functional and structural studies of proteins and a versatile tool for in vitro protein evolution and synthetic biology[14].
3.Experiment Design
Basically, we utilized the cell-free system to express the enzymes which had been modified in 22 different sites. Besides, we added a fluorescet protein, CFP, before the enzyme. And there is a flexible linker, GGGGSGGGGS , between them. So that we could detect the expression of enzymes by detecting expression of the fluorescent protein with a fluorescence readout instrument, for example, a microplate reader. We conceived that with this method we could acquire the best modifications by screening them in a high-throughput way. Then we used the proteins we got to degrade PET.
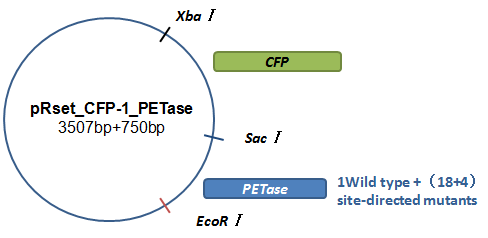
Fig.7.The expression vector in CFPS system
How to characterize the degradation velocity is the main problem in our scheme. We analyzed the experiment consequences in two ways. For the first one, we rendered the enzymes degrade pNPa, a general substituent for the detection of PET. Then we measured the absorbance of pNP in the optical density of 400 nanometers, which is the degrading product of pNPa. For the second one, we detected the absorbance of MHET in the optical density of 260 nanometers, which is the product in the first step of PET degradation.

Fig.8.Steps for integrated expression and activity screening of enzymes
References
[1] Yoshida S, Hiraga K, Takehana T, et al. A bacterium that degrades and assimilates poly (ethylene terephthalate)[J]. Science, 2016, 351(6278): 1196-1199.
[2] Chen S, Tong X, Woodard R W, et al. Identification and characterization of bacterial cutinase[J]. Journal of Biological Chemistry, 2008, 283(38): 25854-25862.
[3] Schwede T, Kopp J, Guex N, Peitsch MC. SWISS-MODEL: an automated protein homologymodeling server. Nucleic Acids Research, 2003, 31 (13): 3381-3385
[4] Yoshida S, Hiraga K, Takehana T, et al. Supplementary Material For A bacterium that degrades and assimilates poly (ethylene terephthalate)
[5] Araújo R, Silva C, O’Neill A, et al. Tailoring cutinase activity towards polyethylene terephthalate and polyamide 6, 6 fibers[J]. Journal of Biotechnology, 2007, 128(4): 849-857.
[6] Silva C, Da S, Silva N, et al. Engineered Thermobifida fusca cutinase with increased activity on polyester substrates[J]. Biotechnology journal, 2011, 6(10): 1230-1239.
[7] Herrero Acero E, Ribitsch D, Dellacher A, et al. Surface engineering of a cutinase from Thermobifida cellulosilytica for improved polyester hydrolysis[J]. Biotechnology and bioengineering, 2013, 110(10): 2581-2590.
[8] Wei R, Oeser T, Schmidt J, et al. Engineered bacterial polyester hydrolases efficiently degrade polyethylene terephthalate due to relieved product inhibition[J]. Biotechnology and bioengineering, 2016.
[9] Ansaldi M, Lepelletier M, Méjean V. Site-specific mutagenesis by using an accurate recombinant polymerase chain reaction method [J]. Analytical biochemistry, 1996, 234(1): 110-111.
[10] Roth C, Wei R, Oeser T, et al. Structural and functional studies on a thermostable polyethylene terephthalate degrading hydrolase from Thermobifida fusca[J]. Applied microbiology and biotechnology, 2014, 98(18): 7815-7823.
[11]Gabriel Rosenblum and Barry S. Cooperman. Engine out of the Chassis: Cell-Free Protein Synthesis and its Uses. FEBS Lett. 2014 January 21; 588(2): 261–268. doi:10.1016/j.febslet.2013.10.016.
[12]Aarthi Chandrasekaran and Anup K. Singh. One-Pot, Microscale Cell-Free Enzyme Expression and Screening. DOI 10.1007/978-1-62703-782-2 Springer New York Heidelberg Dordrecht London.
[13] C. Eric Hodgman, Michael C. Jewett. Cell-free synthetic biology: Thinking outside the cell. Metabolic Engineering 14 (2012) 261–269. doi:10.1016/j.ymben.2011.09.002
[14] Shaorong Chong. Overview of Cell-Free Protein Synthesis: Historic Landmarks, Commercial
Systems, and Expanding Applications. Current Protocols in Molecular Biology 16.30.1-16.30.11, October 2014
DOI: 10.1002/0471142727.mb1630s108


