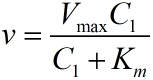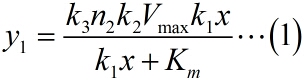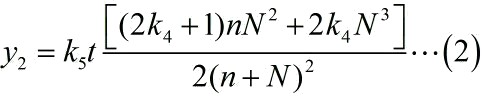Abstract
This model is created to evaluate the effectiveness of initial design, and offers guidelines on how the system can (or must) be improved. (You can go to PROJECT. page to see more.) The core idea is to simulate the process of producing the signal which can be detected, and draw a conclusion by obtaining the relationship between the signal intensity and the concentration of miRNA.
Introduction
We create mathematical models of two aspects of our project, a RCA model and a signal detection model.
The RCA model is based on the Michaelis-Menten equation. We can obtain the relationship between the concentration of miRNA and the number of stem-loop structures through theoretical calculation, and we use our experimental results to calculate the parameters we introduced previously.
In the signal detection model, we combine a probability model with the kinetic equation of enzymatic reaction, so we can obtain the relationship between the number of stem-loop structures and the signal intensity under the premise of adding a certain amount of the fusion proteins of dCas9 and split-HRP fragments.
By integrating the two models, we can theoretically predict the impacts of the molecule number of proteins on the signal to noise ratio and explain the trend of the signal intensity variation against the change of the concentration of miRNA in our wet-lab work.
Assumption and Justification
About model
1. MiRNA is not degraded throughout the reaction process.
2. The two fusion proteins of dCas9 and split-HRP fragments have the same ability to combine with the stem-loop structure, and only when two different proteins next to each other, can they have the ability to catalyze substrate and produce signal.
3. The number of stem-loop structures in each RCA product is equal under a certain reaction time.
4. The enzymatic activity remains unchanged with time under the premise of excessive amount of enzymes or a short-time reaction.
About the data
1. The data we obtain from wet-lab experiment are reliable.
2. All the results are trustworthy in the process of statistical processing and data calculation.
Model
Notations
The RCA model
Firstly, we assume that there is a linear relationship between the concentration of the initiated probes and miRNA as the combination reaction between miRNA and probe occurs spontaneously1 . This can be written as:
Next, RCA is an enzymatic reaction, the reaction equation is written as: iprobe+dNTP→ iprobe-dN+PPi, under the premise of excessive amount of enzymes and dNTPs, the relationship between the concentration of iprobe and the initial speed of RCA can be described by the Michaelis-Menten equation. This can be written as:
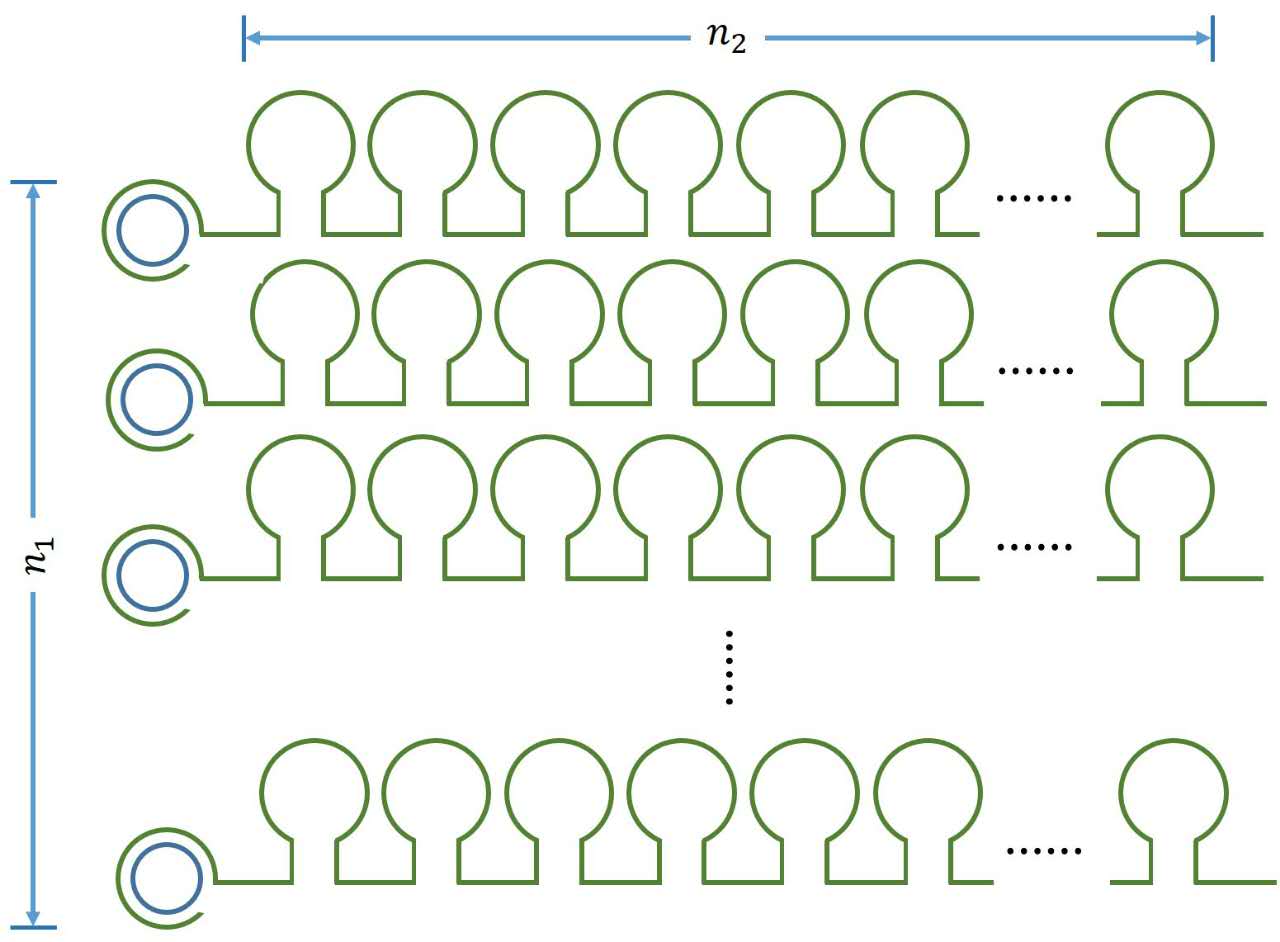
It is obvious that there is a linear relationship between the molecule numbers of iprobe-dN (n1 , reflects the molecule numbers of RCA product) and the initial speed of RCA reaction. Notably, under the premise of excessive amount of enzymes, the extending of each RCA production is related to the reaction time, not the concentration of the iprobe. It is a linear relationship when we assume that the enzymatic activity remains unchanged with time, thus, the number of stem-loop structures (n2 ) in each RCA product is stable under the premise of a certain reaction time.
Figure 1. Schematic diagram
This can be written as:
And then, SYBR Green binds to DNA. The resulting DNA-dye-complex absorbs blue light (λmax = 497nm) and emits green light (λmax = 520nm). Thus, there is a linear relationship between the total amount of stem-loop structures and the fluorescence intensity of DNA-dye-complex. This can be written as:
In summary, the relationship between the concentration of miRNA and the fluorescence intensity of DNA-dye-complex can be written as:
The signal detection model
Firstly, it is obvious that there is a linear relationship between the molecule number of formed intact HRP proteins in the solution (consider as NOISE) and the molecule number of the fusion proteins in the solution. This can be written as:
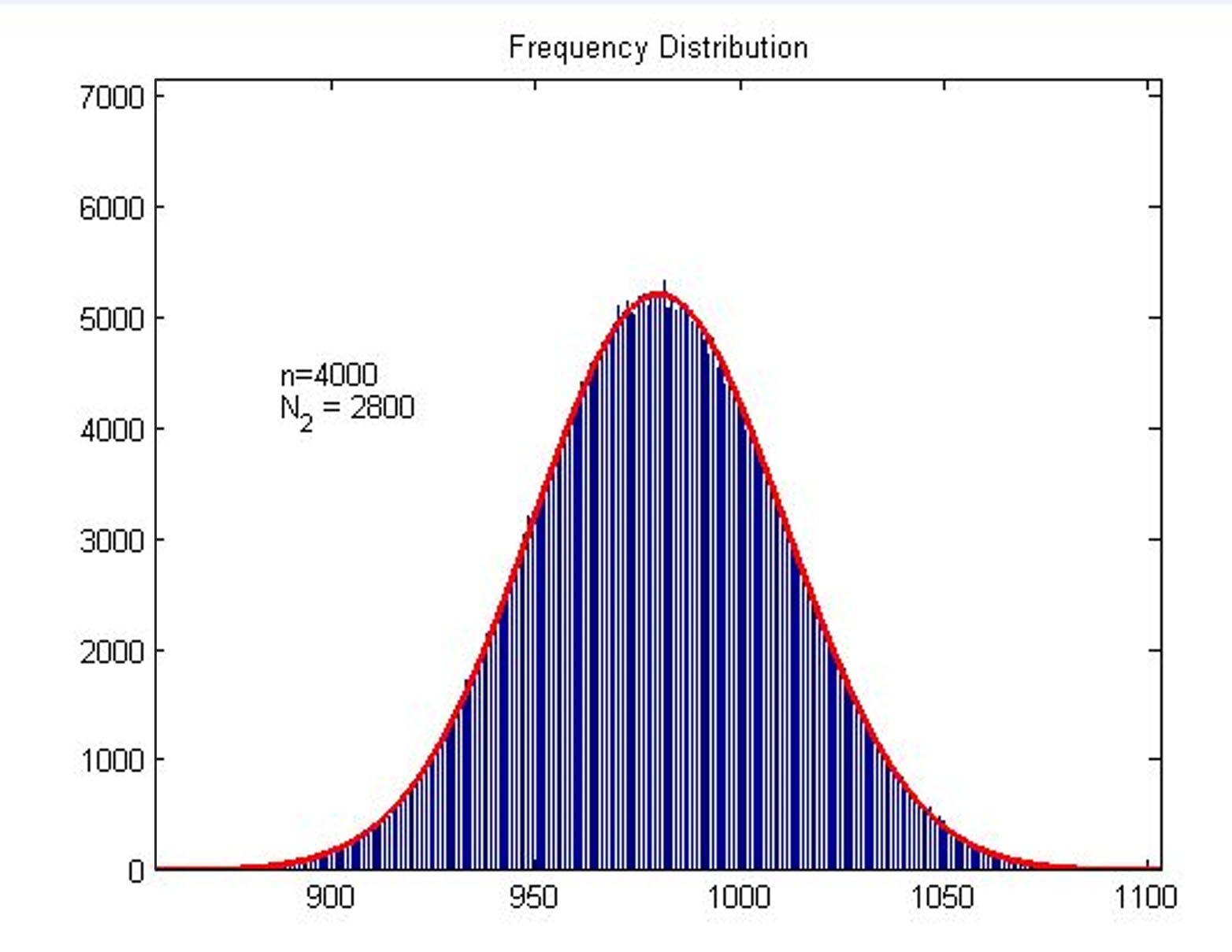
Next, we build a probability model to find the expression of the molecule number of formed intact HRP proteins (fused with dCas9) through binding with stem-loop structure (consider as SIGNAL). It is related to the total amount of stem-loop structures and the molecule number of DNA-bound fusion proteins. We obtain the result by Monte Carlo method.
Figure2. Frequency Distribution Plot of frequency distribution of the molecule number of formed intact HRP proteins through binding with stem-loop structure.
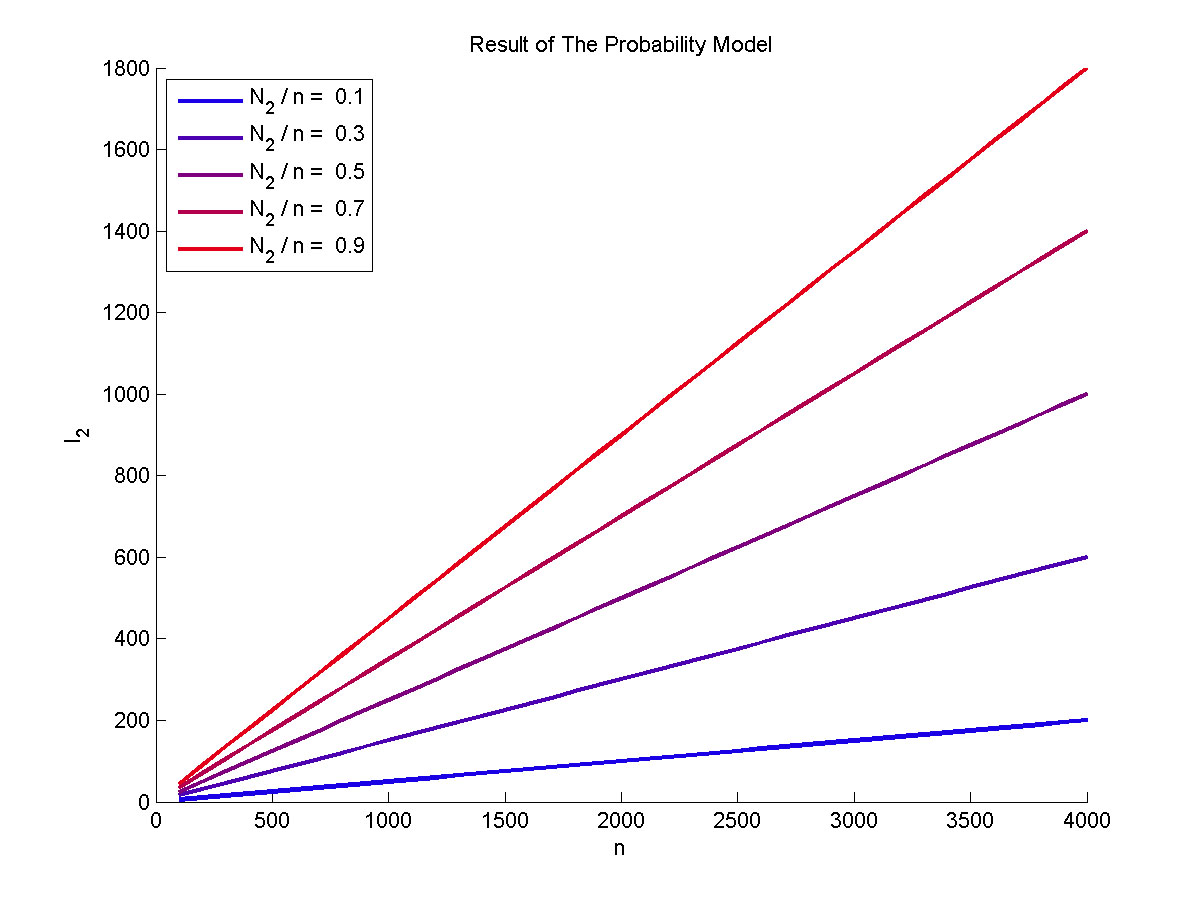
Figure 3. The result of the probability model Plot of the expect value of the molecule number of formed intact proteins against the total amount of stem-loop structures, carried out at 0.1, 0.3, 0.5, 0.7, 0.9 of N2 to n ratio.
The result can be written as:
As for N2, which refers to the molecule number of the fusion proteins binding with stem-loop structure. The equation is formulated based on the limiting case. The equation can be written as:
And then, we can obtain the expressions for ρ and I separately.
Notably, under the premise of excessive substrates, it is a linear relationship between the reaction rate and the concentration of enzymes when we assume that the enzymatic activity remain unchanged with time, thus, we can obtain the expression of the signal intensity at tth time-step. This is written as:
In summary, the relationship between the number of stem-loop structures and SNR under the premise of adding a certain amount of the fusion proteins can be written as:
The relationship between the number of Stem-loop structures and the signal intensity under the premise of adding a certain amount of the fusion proteins can be written as:
The calculation of the constants
To simplify the equation (1), the constants can be integrated. As can be seen from the above table, k1, k2, k3 are constants representing the scale factor, Km and Vmax are characteristic constants of phi 29 DNA polymerase, and n2, which refers to the number of stem-loop structures in each RCA product, is stable under the premise of a certain reaction time. Thus, define two constants, mark as a and b, and then the equation can be simplified as:
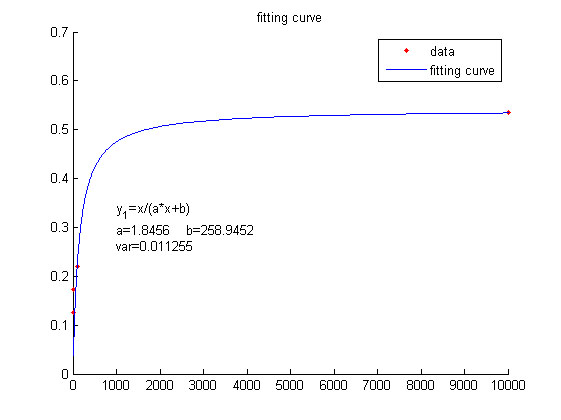
We use this equation to fit the data points obtained through experiments. (Figure 2. (D)in the RESULTS.page) The fitting curve is shown below.
Figure 4. Fitting curve Fitting curve of fluorescence intensity of DNA-dye-complex (RFU) against the concentrations of miRNA (pM).
With regard to the equation (2), we set the parameters based on the experiments as follows: k3 =8.35*10-16 (n=y1/k3); k4 =0.038; k5t=5.9*10-12.
Results and Analysis
By integrating the two models, we can obtain the relationships between SNR, the signal intensity respectively and the concentration of miRNA under different addition amount of fusion proteins.
The results are shown below.
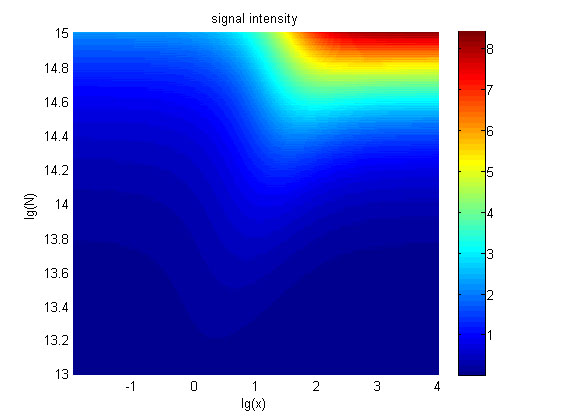
Figure 5. The signal intensity (OD450) Three-dimensional map of signal intensity (OD450) against miRNA concentration(pM) and additional amount of fusion proteins.
There exists a monotonous relation between the concentration of miRNA and the signal intensity when the value of the molecule number of the fusion proteins is relatively large. While the relationship does not hold when the value of the molecule number of the fusion proteins is relatively small. And the signal intensity increases as the value of the molecule number of the fusion proteins increases.
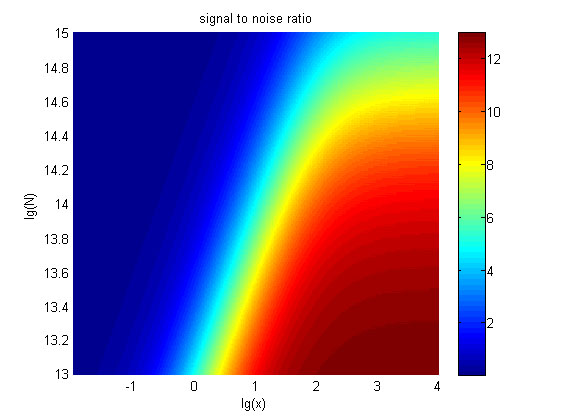
Figure 6. The result of SNR Three-dimensional map of signal to noise ratio against miRNA concentration (pM) and additional amount of fusion proteins.
The signal-to-noise ratio decreases as the value of the molecule number of fusion proteins increases.
Test of model
Our model predicts the relationship between the signal intensity and the concentration of miRNA under the premise of adding a certain amount of the fusion proteins of dCas9 and split-HRP fragments. So we can test if our model is reasonable by comparing the curves predicted by the model with those obtained from the experiment. The results are shown below.
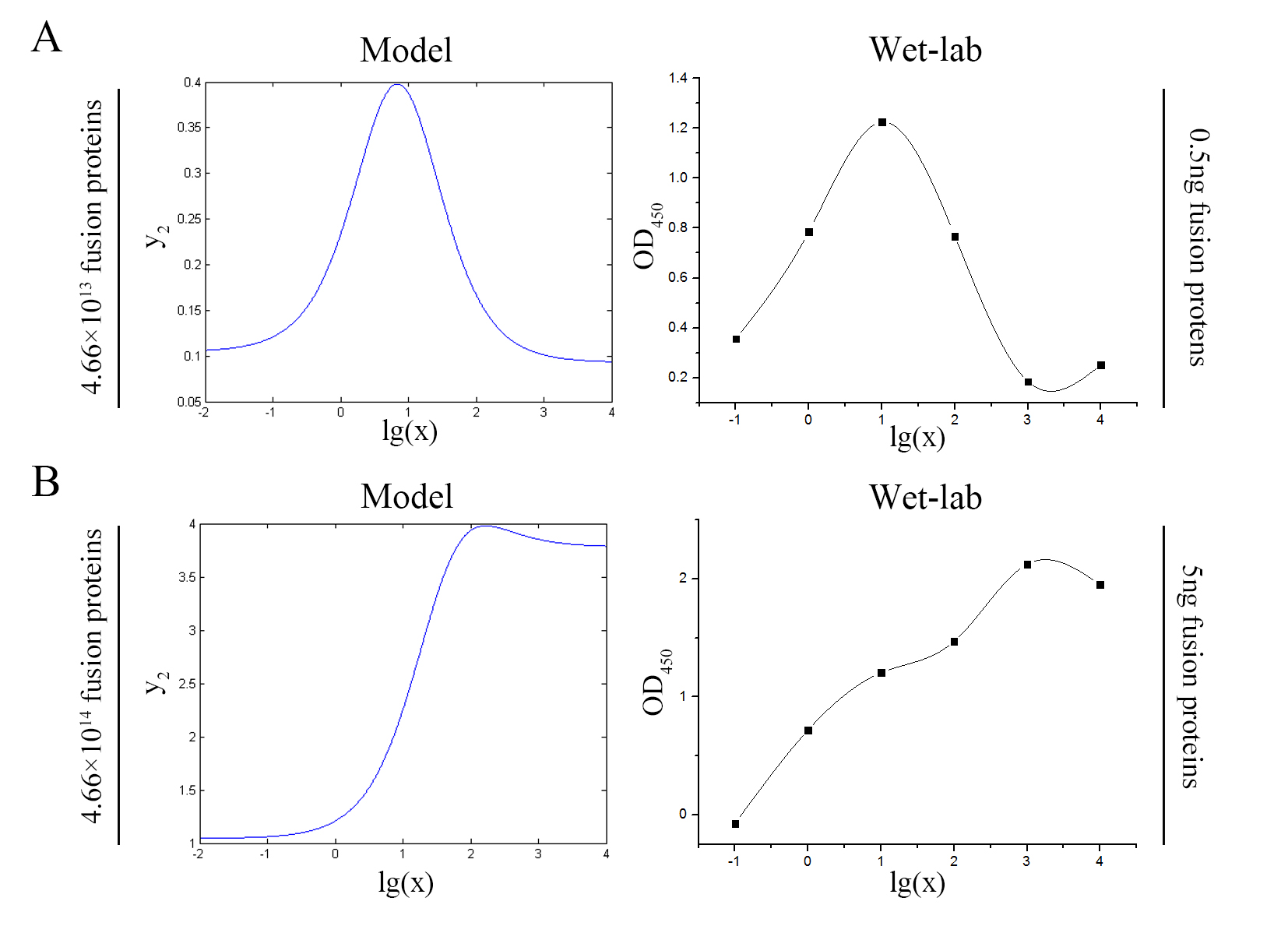
Figure 7. Test of model Plot of the signal intensity (OD450) against miRNA concentration (pM) predicted by the model and experimented by wet-lab. (A) The value of the molecule number of each fusion protein added to the system is set to be 4.66e+13. The mass of each fusion protein added to the system in wet-lab is 0.5 ng. (B) The value of the molecule number of each fusion protein added to the system is set to be 4.66e+14. The mass of each fusion protein added to the system in wet-lab is 5 ng.
We can learn from the figure above that the general trend of curves predicted by the model agree well with the experimental results.
Model extension
Furthermore, we consider that it is conducive to signal detection when the value of signal intensity and SNR are both greater than two. We obtain the ideal region through calculation, which is shown as below.
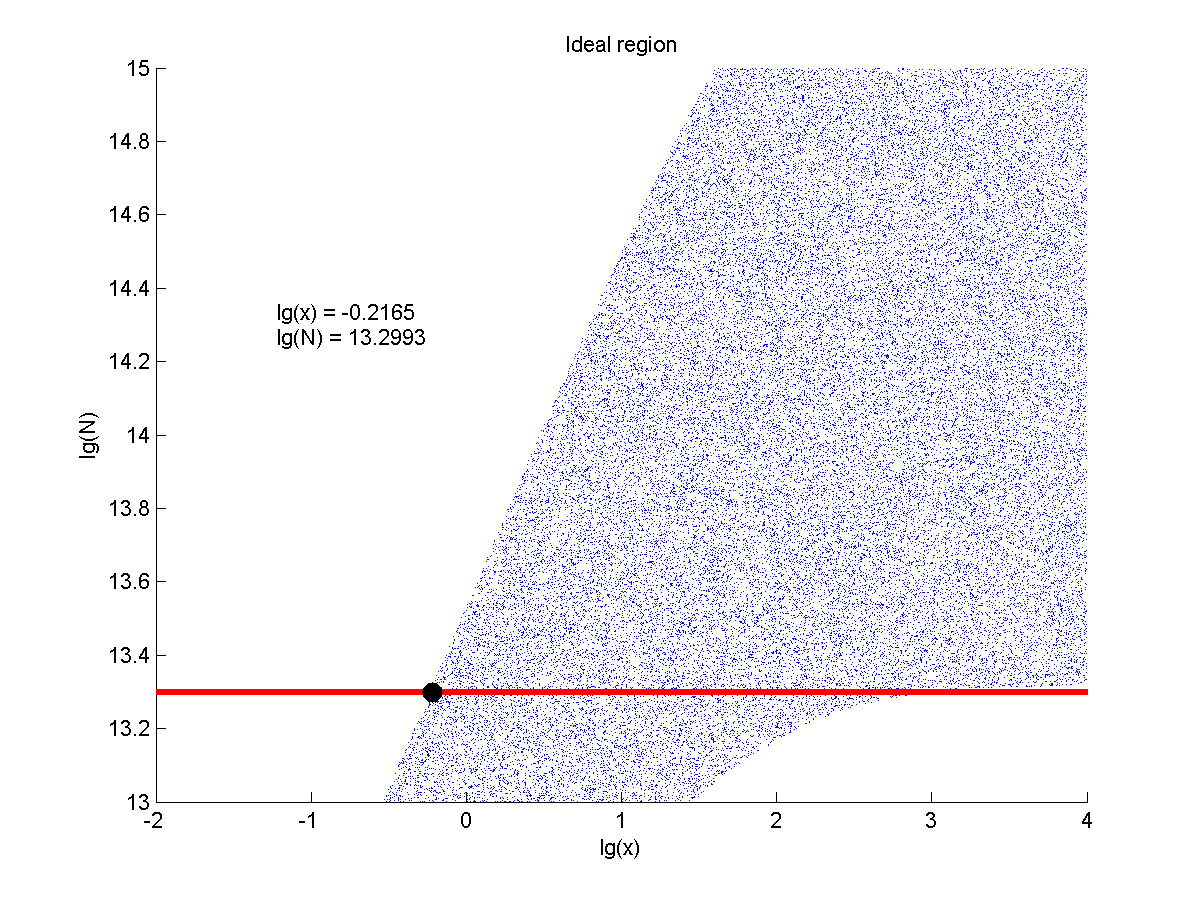
Figure 8. Ideal region Region of qualified logarithm of concentrations of miRNA (pM) and logarithm of the additional amount of fusion proteins.
When the value of N is set to e+13.3, the range of the value of x contained in the ideal region is a maximum, which means when the value of the molecule number of the fusion proteins is e+13.3, the concentration range of miRNA that can be detected is maximum.
Conclusion
Based on our simulation, we came to the conclusion that:
The general trend of curves predicted by the model are in good agreement with the experimental results, thus indicating that our model is reasonable.
The value of the molecule number of the fusion protein of dCas9 and split-HRP fragments not only affects the signal-to-noise ratio, but also the signal intensity. So we need to weigh its impact on both to select the optimal solution.
The concentration range of miRNA that can be detected is a maximum when we set the value of the molecule number of the fusion proteins to be e+13.3, building on which, our wet-lab protocol could be optimized in our future work.
Reference
1 Deng, R. et al.Toehold-initiated rolling circleamplification for visualizing individual microRNAs in situ in single cells. Angew Chem Int Ed Engl 53, 2389-2393,doi:10.1002/anie.201309388 (2014).