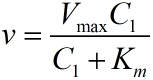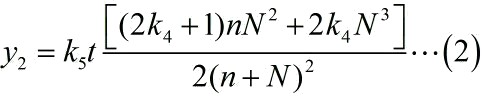Zhuchushu13 (Talk | contribs) |
Zhuchushu13 (Talk | contribs) |
||
| (12 intermediate revisions by 2 users not shown) | |||
| Line 388: | Line 388: | ||
model is created to evaluate the effectiveness of initial design, and offers | model is created to evaluate the effectiveness of initial design, and offers | ||
| − | guidelines | + | guidelines on how the system can (or must) be improved. (You can go to <a href="/Team:NUDT_CHINA/Design">PROJECT. page</a> to see more.) The core idea is to simulate the process of producing the signal which can be detected, and draw a conclusion by obtaining the relationship between the signal intensity and the concentration of miRNA. |
</span> | </span> | ||
</p> | </p> | ||
| Line 446: | Line 446: | ||
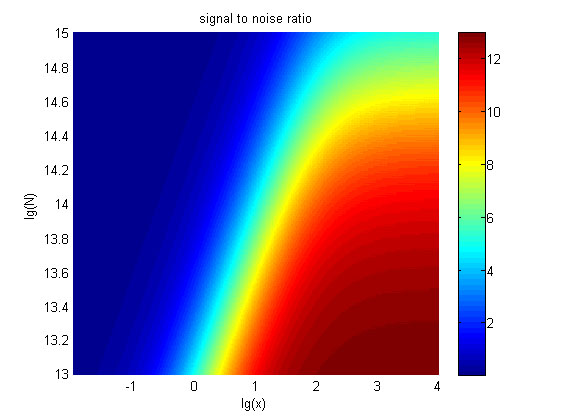
integrating the two models, we can theoretically predict the impacts of the | integrating the two models, we can theoretically predict the impacts of the | ||
| − | molecule number of proteins on the signal to noise ratio and | + | molecule number of proteins on the signal to noise ratio and explain the trend |
| − | of the signal intensity | + | of the signal intensity variation against the change of the concentration of miRNA in our |
wet-lab work. | wet-lab work. | ||
| Line 465: | Line 465: | ||
<!--标题--> | <!--标题--> | ||
<h3> | <h3> | ||
| − | <span><span style="color:#7f1015"> About model </span></span | + | <span><span style="color:#7f1015"> About model </span></span> |
</h3> | </h3> | ||
| Line 471: | Line 471: | ||
<!--正文--> | <!--正文--> | ||
<p style="text-indent:22pt;"> | <p style="text-indent:22pt;"> | ||
| − | <span style="line-height:2;font-family:Perpetua;font-size:18px;"> | + | <span style="line-height:2;font-family:Perpetua;font-size:18px;">1. MiRNA is not degraded throughout the reaction process.</span> |
| − | + | ||
| − | + | ||
</p> | </p> | ||
</br> | </br> | ||
| Line 480: | Line 478: | ||
<!--正文--> | <!--正文--> | ||
<p style="text-indent:22pt;"> | <p style="text-indent:22pt;"> | ||
| − | <span style="line-height:2;font-family:Perpetua;font-size:18px;"> | + | <span style="line-height:2;font-family:Perpetua;font-size:18px;">2. The two fusion proteins of dCas9 and split-HRP fragments have the |
| − | + | ||
| − | + | ||
same ability to combine with the stem-loop structure, and only when two | same ability to combine with the stem-loop structure, and only when two | ||
| Line 495: | Line 491: | ||
<!--正文--> | <!--正文--> | ||
<p style="text-indent:22pt;"> | <p style="text-indent:22pt;"> | ||
| − | <span style="line-height:2;font-family:Perpetua;font-size:18px;"> | + | <span style="line-height:2;font-family:Perpetua;font-size:18px;">3. The number of stem-loop structures in each RCA product is equal |
| − | + | ||
| − | + | ||
under a certain reaction time. </span> | under a certain reaction time. </span> | ||
| Line 506: | Line 500: | ||
<!--正文--> | <!--正文--> | ||
<p style="text-indent:22pt;"> | <p style="text-indent:22pt;"> | ||
| − | <span style="line-height:2;font-family:Perpetua;font-size:18px;"> | + | <span style="line-height:2;font-family:Perpetua;font-size:18px;">4. The enzymatic activity remains unchanged with time under the premise |
| − | + | ||
| − | + | ||
of excessive amount of enzymes or a short-time reaction.</span> | of excessive amount of enzymes or a short-time reaction.</span> | ||
| Line 518: | Line 510: | ||
<!--标题--> | <!--标题--> | ||
<h3> | <h3> | ||
| − | <span><span style="color:#7f1015"> About the data </span></span | + | <span><span style="color:#7f1015"> About the data </span></span> |
| − | + | ||
| − | + | ||
</h3> | </h3> | ||
| Line 526: | Line 516: | ||
<!--正文--> | <!--正文--> | ||
<p style="text-indent:22pt;"> | <p style="text-indent:22pt;"> | ||
| − | <span style="line-height:2;font-family:Perpetua;font-size:18px;"> | + | <span style="line-height:2;font-family:Perpetua;font-size:18px;">1. The data we obtain from wet-lab experiment are reliable.</span> |
| − | + | ||
| − | + | ||
</p> | </p> | ||
</br> | </br> | ||
| Line 535: | Line 523: | ||
<!--正文--> | <!--正文--> | ||
<p style="text-indent:22pt;"> | <p style="text-indent:22pt;"> | ||
| − | <span style="line-height:2;font-family:Perpetua;font-size:18px;"> | + | <span style="line-height:2;font-family:Perpetua;font-size:18px;">2. All the results are trustworthy in the process of statistical |
| − | + | ||
| − | + | ||
processing and data calculation.</span> | processing and data calculation.</span> | ||
| Line 551: | Line 537: | ||
<!--标题--> | <!--标题--> | ||
<h3> | <h3> | ||
| − | <span><span style="color:#7f1015">Notations </span></span | + | <span><span style="color:#7f1015">Notations </span></span> |
</h3> | </h3> | ||
| Line 557: | Line 543: | ||
<!--表格--> | <!--表格--> | ||
| − | <table class="MsoTableGrid" border="1" cellspacing="0" cellpadding="0" | + | <p> |
| − | + | <table class="MsoTableGrid" border="1" cellspacing="0" cellpadding="0" style="border:none;"> | |
| − | style="border:none;"> | + | <tbody> |
| − | + | <tr> | |
| − | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <span style="font-family:"">Symbol </span> | |
| − | windowtext 1.0pt;"> | + | </p> |
| − | + | </td> | |
| − | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <span style="font-family:"">Definition </span> | |
| − | + | </p> | |
| − | + | </td> | |
| − | windowtext 1.0pt;"> | + | </tr> |
| − | + | <tr> | |
| − | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <i><span style="font-family:"">x</span></i> | |
| − | + | </p> | |
| − | + | </td> | |
| − | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | windowtext 1.0pt;"> | + | <a name="OLE_LINK7"></a><a name="OLE_LINK6"></a><span style="font-family:"">The |
| − | + | concentration of</span><span style="font-family:""> miRNA (pM)</span> | |
| − | + | </p> | |
| − | + | </td> | |
| − | + | </tr> | |
| − | + | <tr> | |
| − | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | windowtext 1.0pt;"> | + | <p class="MsoNormal"> |
| − | + | <i><span style="font-family:"">C<sub>1</sub></span></i> | |
| − | + | </p> | |
| − | + | </td> | |
| − | name="OLE_LINK6"></a><span>The concentration of</span><span> miRNA (pM)</span> | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> |
| − | + | <p class="MsoNormal"> | |
| − | + | <span style="font-family:"">The | |
| − | + | concentration of initiated probe (Abbreviated to iprobe)</span> | |
| − | + | </p> | |
| − | + | </td> | |
| − | + | </tr> | |
| − | windowtext 1.0pt;"> | + | <tr> |
| − | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <i><span style="font-family:"">k<sub>1</sub></span></i> | |
| − | + | </p> | |
| − | + | </td> | |
| − | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | windowtext 1.0pt;"> | + | <p class="MsoNormal"> |
| − | + | <a name="OLE_LINK8"></a><span style="font-family:"">A constant representing the scale factor</span><span style="font-family:""></span> | |
| − | + | </p> | |
| − | + | </td> | |
| − | probe (Abbreviated | + | </tr> |
| − | + | <tr> | |
| − | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <i><span style="font-family:"">K<sub>m </sub></span></i> | |
| − | + | </p> | |
| − | + | </td> | |
| − | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | windowtext 1.0pt;"> | + | <p class="MsoNormal"> |
| − | + | <span style="font-family:"">One | |
| − | + | of the characteristic constants of phi29 DNA polymerase</span> | |
| − | + | </p> | |
| − | + | </td> | |
| − | + | </tr> | |
| − | + | <tr> | |
| − | windowtext 1.0pt;"> | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> |
| − | + | <p class="MsoNormal"> | |
| − | + | <i><span style="font-family:"">V<sub>max</sub></span></i> | |
| − | + | </p> | |
| − | representing | + | </td> |
| − | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <span style="font-family:"">One | |
| − | + | of the characteristic constants of phi29 DNA polymerase</span> | |
| − | + | </p> | |
| − | + | </td> | |
| − | + | </tr> | |
| − | windowtext 1.0pt;"> | + | <tr> |
| − | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <i><span style="font-family:"">k<sub>2</sub></span></i> | |
| − | + | </p> | |
| − | + | </td> | |
| − | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | windowtext 1.0pt;"> | + | <p class="MsoNormal"> |
| − | + | <span style="font-family:"">A | |
| − | + | constant representing the scale factor</span> | |
| − | + | </p> | |
| − | constants of | + | </td> |
| − | + | </tr> | |
| − | + | <tr> | |
| − | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <i><span style="font-family:"">V</span></i> | |
| − | + | </p> | |
| − | + | </td> | |
| − | windowtext 1.0pt;"> | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> |
| − | + | <p class="MsoNormal"> | |
| − | + | <span style="font-family:"">The initial | |
| − | + | speed of RCA </span> | |
| − | + | </p> | |
| − | + | </td> | |
| − | + | </tr> | |
| − | windowtext 1.0pt;"> | + | <tr> |
| − | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <a name="OLE_LINK9"></a><i><span style="font-family:"">n<sub>1</sub></span></i><i><sub><span style="font-family:""></span></sub></i> | |
| − | constants of phi29 | + | </p> |
| − | + | </td> | |
| − | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <span style="font-family:"">The | |
| − | + | moles of RCA product</span> | |
| − | + | </p> | |
| − | + | </td> | |
| − | windowtext 1.0pt;"> | + | </tr> |
| − | + | <tr> | |
| − | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <i><span style="font-family:"">n<sub>2</sub></span></i> | |
| − | + | </p> | |
| − | + | </td> | |
| − | windowtext 1.0pt;"> | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> |
| − | + | <p class="MsoNormal"> | |
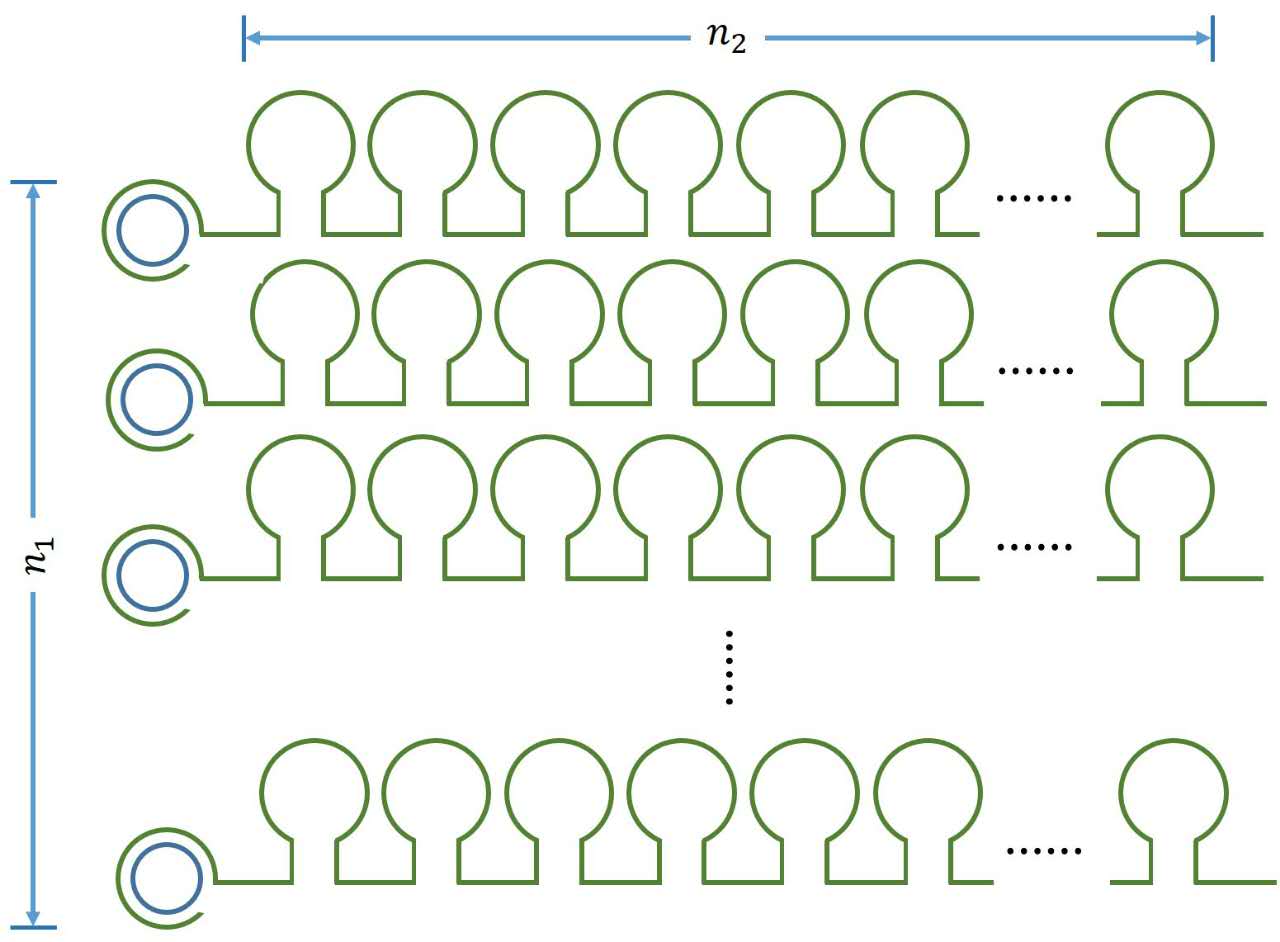
| − | + | <a name="OLE_LINK10"></a><span style="font-family:"">The number of stem-loop structures in each RCA product</span><span style="font-family:""></span> | |
| − | + | </p> | |
| − | factor</span> | + | </td> |
| − | + | </tr> | |
| − | + | <tr> | |
| − | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <i><span style="font-family:"">n</span></i> | |
| − | + | </p> | |
| − | windowtext 1.0pt;"> | + | </td> |
| − | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <span style="font-family:"">The total | |
| − | + | amount of <a name="OLE_LINK23"></a><a name="OLE_LINK22"></a>stem-loop structures</span> | |
| − | + | </p> | |
| − | + | </td> | |
| − | windowtext 1.0pt;"> | + | </tr> |
| − | + | <tr> | |
| − | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <i><span style="font-family:"">N</span></i> | |
| − | + | </p> | |
| − | + | </td> | |
| − | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | windowtext 1.0pt;"> | + | <a name="OLE_LINK41"></a><a name="OLE_LINK16"></a><span style="font-family:"">The |
| − | + | molecule number of the fusion protein of dCas9 and split-HRP fragments</span><span style="font-family:""></span> | |
| − | + | </p> | |
| − | + | </td> | |
| − | name="OLE_LINK9"></a><i><span>n<sub>1</sub></span></i><i><sub><span></span></sub | + | </tr> |
| − | + | <tr> | |
| − | ></i> | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> |
| − | + | <p class="MsoNormal"> | |
| − | + | <i><span style="font-family:"">k<sub>3</sub></span></i> | |
| − | + | </p> | |
| − | + | </td> | |
| − | windowtext 1.0pt;"> | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> |
| − | + | <p class="MsoNormal"> | |
| − | + | <span style="font-family:"">A | |
| − | + | constant representing the scale factor</span> | |
| − | + | </p> | |
| − | + | </td> | |
| − | + | </tr> | |
| − | + | <tr> | |
| − | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | windowtext 1.0pt;"> | + | <p class="MsoNormal"> |
| − | + | <i><span style="font-family:"">y<sub>1</sub></span></i> | |
| − | + | </p> | |
| − | + | </td> | |
| − | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <span style="font-family:"">The fluorescence | |
| − | windowtext 1.0pt;"> | + | intensity of DNA-dye-complex (RFU)</span> |
| − | + | </p> | |
| − | + | </td> | |
| − | + | </tr> | |
| − | number of | + | <tr> |
| − | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <i><span style="font-family:"">N<sub>1</sub></span></i> | |
| − | + | </p> | |
| − | + | </td> | |
| − | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | windowtext 1.0pt;"> | + | <span style="font-family:"">The |
| − | + | molecule number of the fusion protein of dCas9 and split-HRP fragments in the | |
| − | + | solution</span> | |
| − | + | </p> | |
| − | + | </td> | |
| − | + | </tr> | |
| − | + | <tr> | |
| − | windowtext 1.0pt;"> | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> |
| − | + | <p class="MsoNormal"> | |
| − | + | <i><span style="font-family:"">N<sub>2</sub></span></i> | |
| − | + | </p> | |
| − | name="OLE_LINK23"></a><a name="OLE_LINK22"></a>stem-loop structures</span> | + | </td> |
| − | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <a name="OLE_LINK45"></a><span style="font-family:"">The molecule number of the fusion protein of dCas9 and | |
| − | + | split-HRP fragments binding with stem-loop structure</span><span style="font-family:""></span> | |
| − | + | </p> | |
| − | + | </td> | |
| − | windowtext 1.0pt;"> | + | </tr> |
| − | + | <tr> | |
| − | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <i><span style="font-family:"">k<sub>4</sub></span></i> | |
| − | + | </p> | |
| − | + | </td> | |
| − | windowtext 1.0pt;"> | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> |
| − | + | <p class="MsoNormal"> | |
| − | + | <span style="font-family:"">A | |
| − | + | constant representing the scale factor</span> | |
| − | name="OLE_LINK16"></a><span>The molecule number of the | + | </p> |
| − | + | </td> | |
| − | + | </tr> | |
| − | + | <tr> | |
| − | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <i><span style="font-family:"">k<sub>5</sub></span></i> | |
| − | + | </p> | |
| − | windowtext 1.0pt;"> | + | </td> |
| − | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <span style="font-family:"">A | |
| − | + | constant representing the scale factor</span> | |
| − | + | </p> | |
| − | + | </td> | |
| − | windowtext 1.0pt;"> | + | </tr> |
| − | + | <tr> | |
| − | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | factor</span> | + | <i><span style="font-family:"">ρ</span></i><i><span style="font-family:""></span></i> |
| − | + | </p> | |
| − | + | </td> | |
| − | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <span style="font-family:"">Signal | |
| − | + | to noise ratio(Abbreviated to SNR)</span> | |
| − | windowtext 1.0pt;"> | + | </p> |
| − | + | </td> | |
| − | + | </tr> | |
| − | + | <tr> | |
| − | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <i><span style="font-family:"">I</span></i> | |
| − | windowtext 1.0pt;"> | + | </p> |
| − | + | </td> | |
| − | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | DNA-dye-complex (RFU)</span> | + | <p class="MsoNormal"> |
| − | + | <span style="font-family:"">The | |
| − | + | molecule number of formed intact HRP proteins </span> | |
| − | + | </p> | |
| − | + | </td> | |
| − | + | </tr> | |
| − | + | <tr> | |
| − | windowtext 1.0pt;"> | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> |
| − | + | <p class="MsoNormal"> | |
| − | + | <i><span style="font-family:"">I<sub>1</sub></span></i> | |
| − | + | </p> | |
| − | + | </td> | |
| − | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | windowtext 1.0pt;"> | + | <span style="font-family:"">The |
| − | + | molecule number of formed intact HRP proteins in the solution</span> | |
| − | + | </p> | |
| − | + | </td> | |
| − | protein | + | </tr> |
| − | + | <tr> | |
| − | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <i><span style="font-family:"">I<sub>2</sub></span></i> | |
| − | + | </p> | |
| − | + | </td> | |
| − | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | windowtext 1.0pt;"> | + | <p class="MsoNormal"> |
| − | + | <a name="OLE_LINK43"></a><span style="font-family:"">The molecule number of formed intact HRP proteins through | |
| − | + | binding with stem-loop structure</span><span style="font-family:""></span> | |
| − | + | </p> | |
| − | + | </td> | |
| − | + | </tr> | |
| − | + | <tr> | |
| − | windowtext 1.0pt;"> | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> |
| − | + | <p class="MsoNormal"> | |
| − | + | <i><span style="font-family:"">t</span></i> | |
| − | + | </p> | |
| − | molecule number | + | </td> |
| − | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | + | <span style="font-family:"">Reaction | |
| − | + | time</span> | |
| − | + | </p> | |
| − | + | </td> | |
| − | + | </tr> | |
| − | + | <tr> | |
| − | windowtext 1.0pt;"> | + | <td width="66" valign="top" style="border:solid windowtext 1.0pt;"> |
| − | + | <p class="MsoNormal"> | |
| − | + | <i><span style="font-family:"">y<sub>2</sub></span></i> | |
| − | + | </p> | |
| − | + | </td> | |
| − | + | <td width="487" valign="top" style="border:solid windowtext 1.0pt;"> | |
| − | + | <p class="MsoNormal"> | |
| − | windowtext 1.0pt;"> | + | <span style="font-family:"">The |
| − | + | signal intensity (OD<sub>450</sub>)</span> | |
| − | + | </p> | |
| − | + | </td> | |
| − | factor</span> | + | </tr> |
| − | + | </tbody> | |
| − | + | </table> | |
| − | + | </p> | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | windowtext 1.0pt;"> | + | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | windowtext 1.0pt;"> | + | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | factor</span> | + | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | windowtext 1.0pt;"> | + | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | windowtext 1.0pt;"> | + | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | to SNR)</span> | + | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | windowtext 1.0pt;"> | + | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | windowtext 1.0pt;"> | + | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | intact HRP | + | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | windowtext 1.0pt;"> | + | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | windowtext 1.0pt;"> | + | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | intact HRP | + | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | windowtext 1.0pt;"> | + | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | windowtext 1.0pt;"> | + | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | molecule number | + | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | windowtext 1.0pt;"> | + | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | windowtext 1.0pt;"> | + | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | windowtext 1.0pt;"> | + | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | windowtext 1.0pt;"> | + | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | (OD<sub>450</sub>)</span> | + | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | </table> | + | |
| − | + | ||
| Line 1,020: | Line 864: | ||
<!--标题--> | <!--标题--> | ||
<h3> | <h3> | ||
| − | <span><span style="color:#7f1015"> The RCA model </span></span | + | <span><span style="color:#7f1015"> The RCA model </span></span> |
| − | + | ||
| − | + | ||
</h3> | </h3> | ||
| Line 1,195: | Line 1,037: | ||
<span><span style="color:#7f1015"> The signal detection model | <span><span style="color:#7f1015"> The signal detection model | ||
| − | </span></span | + | </span></span> |
</h3> | </h3> | ||
| Line 1,417: | Line 1,259: | ||
<span><span style="color:#7f1015"> The calculation of the constants | <span><span style="color:#7f1015"> The calculation of the constants | ||
| − | </span></span | + | </span></span> |
</h3> | </h3> | ||
| Line 1,455: | Line 1,297: | ||
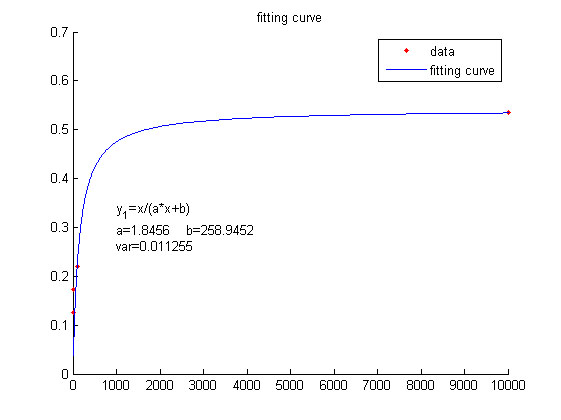
this equation to fit the data points obtained through experiments. (Figure 2. | this equation to fit the data points obtained through experiments. (Figure 2. | ||
| − | (D)in the <a href="/Team:NUDT_CHINA/Results">RESULTS.page</a | + | (D)in the <a href="/Team:NUDT_CHINA/Results">RESULTS.page</a>) The fitting |
curve is shown below.</span> | curve is shown below.</span> | ||
| Line 1,501: | Line 1,343: | ||
<span><span style="color:#7f1015"> Results and Analysis | <span><span style="color:#7f1015"> Results and Analysis | ||
| − | </span></span | + | </span></span> |
</h3> | </h3> | ||
| Line 1,595: | Line 1,437: | ||
<span><span style="color:#7f1015"> Test of model | <span><span style="color:#7f1015"> Test of model | ||
| − | </span></span | + | </span></span> |
</h3> | </h3> | ||
| Line 1,620: | Line 1,462: | ||
</br> | </br> | ||
</html> | </html> | ||
| − | |||
[[File:T--NUDT_CHINA--modelfig7-2.jpg|700px|center]] | [[File:T--NUDT_CHINA--modelfig7-2.jpg|700px|center]] | ||
<html> | <html> | ||
| Line 1,627: | Line 1,468: | ||
<span style="line-height:2;font-family:Perpetua;font- | <span style="line-height:2;font-family:Perpetua;font- | ||
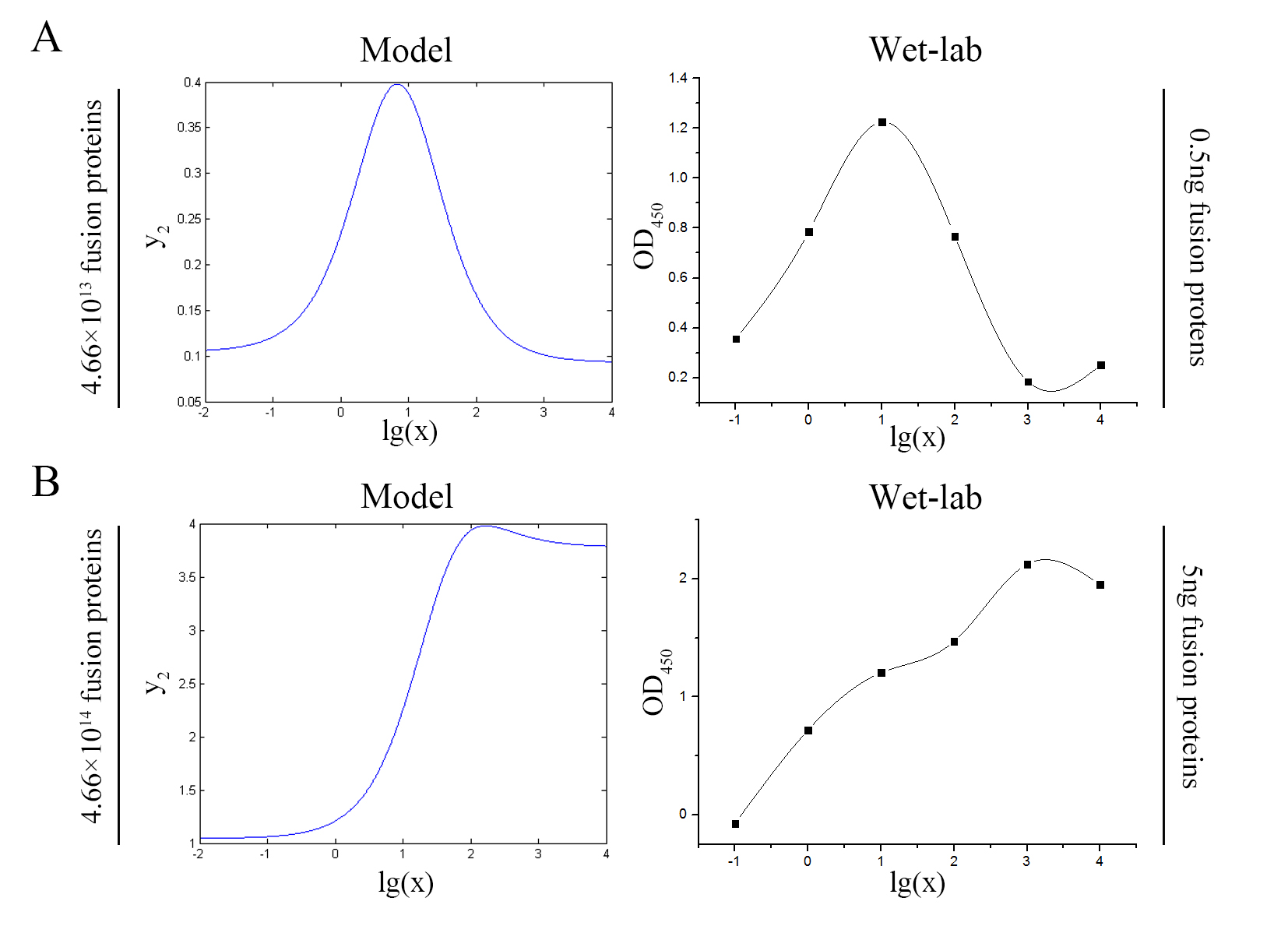
| − | size:18px;"><b>Figure 7. Test of model</b> | + | size:18px;"><b>Figure 7. Test of model</b> Plot of the signal intensity (OD<sub>450</sub>) against miRNA concentration (pM) predicted by the model and experimented by wet-lab. (A) The value of the molecule number of each fusion protein added to the system is set to be 4.66e+13. The mass of each fusion protein added to the system in wet-lab is 0.5 ng. (B) The value of the molecule number of each fusion protein added to the system is set to be 4.66e+14. The mass of each fusion protein added to the system in wet-lab is 5 ng.</span> |
| − | intensity (OD<sub>450</sub>) against miRNA concentration (pM) predicted by the | + | |
| − | model | + | |
| − | 4.66e+13. | + | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | to be 4.66e+14. | + | |
| − | + | ||
| − | + | ||
</p> | </p> | ||
| Line 1,649: | Line 1,479: | ||
<span style="line-height:2;font-family:Perpetua;font-size:18px;">We can learn | <span style="line-height:2;font-family:Perpetua;font-size:18px;">We can learn | ||
| − | from the figure above that the curves predicted by the model | + | from the figure above that the general trend of curves predicted by the model agree well with the experimental results. </span> |
| − | + | ||
| − | + | ||
</p> | </p> | ||
</br> | </br> | ||
| Line 1,661: | Line 1,489: | ||
<span><span style="color:#7f1015"> Model extension | <span><span style="color:#7f1015"> Model extension | ||
| − | </span></span | + | </span></span> |
</h3> | </h3> | ||
| Line 1,704: | Line 1,532: | ||
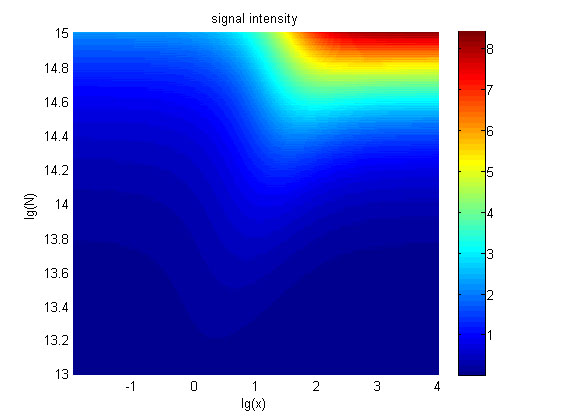
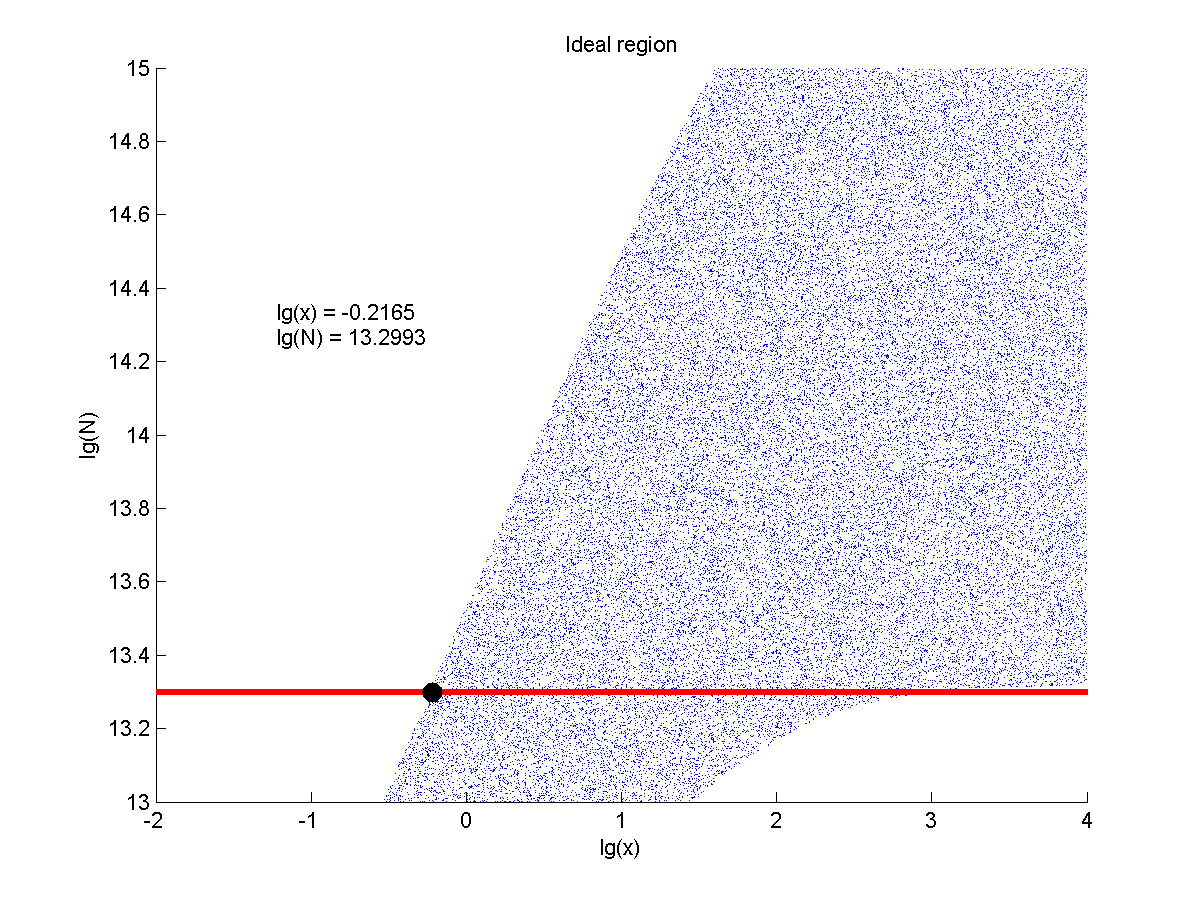
the value of N is set to e+13.3, the range of the value of x contained in the | the value of N is set to e+13.3, the range of the value of x contained in the | ||
| − | ideal region is maximum, which means when the value of the molecule number of | + | ideal region is a maximum, which means when the value of the molecule number of |
the fusion proteins is e+13.3, the concentration range of miRNA that can be | the fusion proteins is e+13.3, the concentration range of miRNA that can be | ||
| Line 1,729: | Line 1,557: | ||
<!--正文--> | <!--正文--> | ||
<p style="text-indent:22pt;"> | <p style="text-indent:22pt;"> | ||
| − | <span style="line-height:2;font-family:Perpetua;font-size:18px;">The | + | <span style="line-height:2;font-family:Perpetua;font-size:18px;">The general trend of curves predicted by the model are in good agreement with the experimental |
| − | + | ||
| − | curves predicted by the model are in good agreement with the experimental | + | |
results, thus indicating that our model is reasonable.</span> | results, thus indicating that our model is reasonable.</span> | ||
| Line 1,756: | Line 1,582: | ||
<span style="line-height:2;font-family:Perpetua;font-size:18px;">The | <span style="line-height:2;font-family:Perpetua;font-size:18px;">The | ||
| − | concentration range of miRNA that can be detected is maximum when we set the | + | concentration range of miRNA that can be detected is a maximum when we set the |
value of the molecule number of the fusion proteins to be e+13.3, building on | value of the molecule number of the fusion proteins to be e+13.3, building on | ||
Latest revision as of 13:34, 19 October 2016
TOP