Zhuchushu13 (Talk | contribs) |
|||
| Line 353: | Line 353: | ||
<div style="text-align:right;"><h5> | <div style="text-align:right;"><h5> | ||
<a style="color:rgb(10,31,84);" href="/Team:NUDT_CHINA">HomePage</a> • | <a style="color:rgb(10,31,84);" href="/Team:NUDT_CHINA">HomePage</a> • | ||
| − | <!-- | + | <!-- 修改这里!! -->TEAM • |
| − | <a style="color:rgb(10,31,84);" href="/Team:NUDT_CHINA/Collaborations"><!-- | + | <a style="color:rgb(10,31,84);" href="/Team:NUDT_CHINA/Collaborations"><!-- 修改这里!! -->Collaborations</a> |
</h5><hr style="width:40%;margin-left:60%;border-top:1px solid rgb(10,31,84);" /></div> | </h5><hr style="width:40%;margin-left:60%;border-top:1px solid rgb(10,31,84);" /></div> | ||
| Line 361: | Line 361: | ||
| − | + | <!--标题--> | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
<h2> | <h2> | ||
| − | <span><span style="color:#7f1015"> | + | <span><span style="color:#7f1015">Collaboration with NJU-China</span></span><hr /> |
| − | + | ||
| − | /> | + | |
</h2> | </h2> | ||
<!--正文--> | <!--正文--> | ||
<p style="text-indent:22pt;"> | <p style="text-indent:22pt;"> | ||
| − | <span style="line-height:2;font-family:Perpetua;font-size:18px;"> | + | <span style="line-height:2;font-family:Perpetua;font-size:18px;">NJU-China performed a Cas9-digest assay to help us validating the effectiveness of their sgRNAs, since our team met some trouble in assessing the efficiency of the single guide RNAs we designed for our further experiments. Also, they helped us to optimize our own protocol for in vitro expression of sgRNAs, so that our project could be performed more smoothly. </span> |
| + | </p> | ||
| + | </br> | ||
| − | + | <!--正文--> | |
| − | + | <p style="text-indent:22pt;"> | |
| − | + | <span style="line-height:2;font-family:Perpetua;font-size:18px;">In return, we performed TEM imaging of exosomes carrying KRAS siRNA and expressing iRGD peptide on their membrane for NJU-China, as well as nanoparticle tracking analysis (NTA) to help them to ascertain the relationship between protein concentration and exosomes quantity.</span> | |
| − | + | ||
| − | + | ||
</p> | </p> | ||
</br> | </br> | ||
| + | |||
| + | |||
| + | <!--标题--> | ||
| + | <h3> | ||
| + | <span><span style="color:#7f1015">Collaboration with NJU-China</span></span><hr /> | ||
| + | </h3> | ||
<!--正文--> | <!--正文--> | ||
<p style="text-indent:22pt;"> | <p style="text-indent:22pt;"> | ||
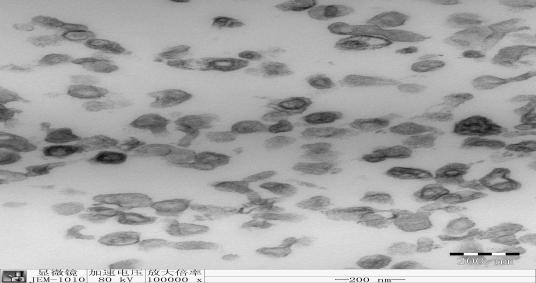
| − | <span style="line-height:2;font-family:Perpetua;font-size:18px;"> | + | <span style="line-height:2;font-family:Perpetua;font-size:18px;">After co-transfection of the two plasmids mentioned above, we performed a transmission electron microscopy (TEM) to characterize the iRGD-modified siRNA-loaded exosomes. The TEM image showed that the exosomes presented normal morphological characteristics after outside modification and siRNA loading, with a diameter of approximately 200 nanometer and a double-layer membrane. These characteristics indicate that the exosome properties won’t be affected by our modifications. </span> |
| + | </p> | ||
| + | </br> | ||
| − | + | <!--插入图片--> | |
| + | </br> | ||
| + | </html> | ||
| + | [[File:T--NUDT_CHINA--collaborationsfig1.jpg|700px|center]] | ||
| + | <html> | ||
| + | </br> | ||
| − | + | <!--图注--> | |
| + | <p> | ||
| + | <span style="line-height:2;font-family:Perpetua;font-size:16px;text-align:center;font-weight:bold">Figure1.TEM image of iRGD-modified exosomes packaging KRAS siRNA</span> | ||
</p> | </p> | ||
</br> | </br> | ||
| + | |||
| + | <!--标题--> | ||
| + | <h3> | ||
| + | <span><span style="color:#7f1015">Nanoparticle tracking analysis to ascertain the relationship between protein concentration and exosomes quantity</span></span><hr /> | ||
| + | </h3> | ||
<!--正文--> | <!--正文--> | ||
<p style="text-indent:22pt;"> | <p style="text-indent:22pt;"> | ||
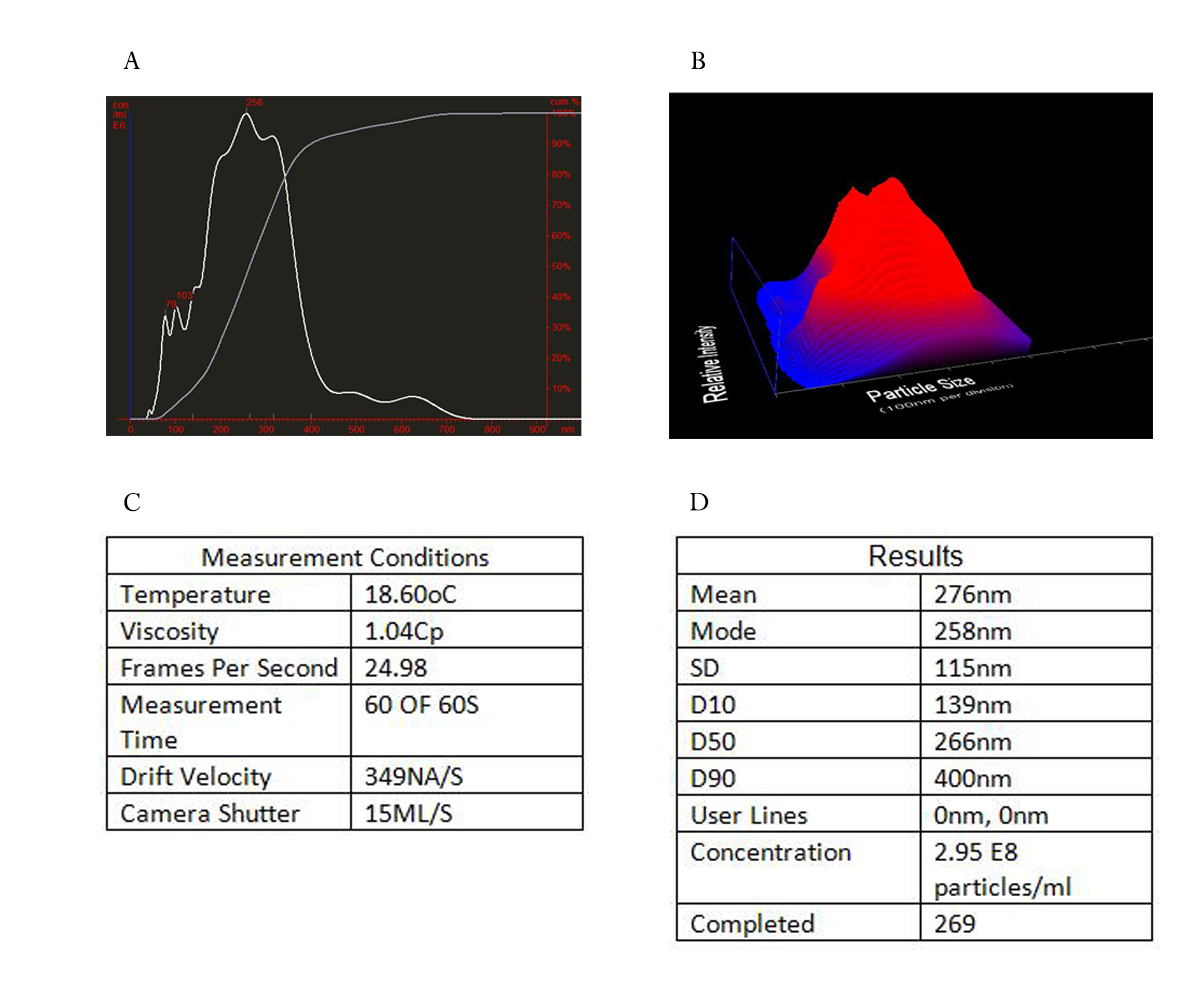
| − | <span style="line-height:2;font-family:Perpetua;font-size:18px;">We | + | <span style="line-height:2;font-family:Perpetua;font-size:18px;">We then performed nanoparticle tracking analysis (NTA) for a further evaluation of the quantity and size of secreted exosomes. The use of Nanosight enabled quantification and size determination of the extracellular vesicles, as nanoparticles can be automatically tracked and sized based on Brownian motion and the diffusion coefficient. The size of exosomes attained ranged around 270nm. Basing the particle size and relative intensity, we also created a 3D plot for a visual explanation. Under measurement condition listed, the exosomes extracted from HEK293 cells were assayed for 2.95 E8 particles each milliliter. Then the relationship between particle number and protein was determined that exosomes in 1 ng protein were equivalent to 6277.95 particles, according to the dilution multiple (24) and protein concentration (1127.756 ng/ul) we have tested. All the data collected help us decide the transfection dosage of siRNA and dosing of treatment prepared for animal experiment.</span> |
| + | </p> | ||
| + | </br> | ||
| − | + | <!--插入图片--> | |
| + | </br> | ||
| + | </html> | ||
| + | [[File:T--NUDT_CHINA--collaborationsfig2.jpg|700px|center]] | ||
| + | <html> | ||
| + | </br> | ||
| − | + | <!--图注--> | |
| + | <p> | ||
| + | <span style="line-height:2;font-family:Perpetua;font-size:16px;font-weight:bold">Figure 2. Nanoparticle tracking analysis (NTA) for characterization of secreted exosomes.</span> | ||
| − | |||
| − | + | <!--图注--> | |
| + | |||
| + | <span style="line-height:2;font-family:Perpetua;font-size:16px;">(a) Concentration of different particle sizes of exosomes. (b) 3D plot of particle size and relative intensity. (c) Experiment condition for our measurement. (d) Results attained after measurement of exosomes.</span> | ||
</p> | </p> | ||
</br> | </br> | ||
| + | |||
| + | <!--插入图片--> | ||
| + | </br> | ||
| + | </html> | ||
| + | [[File:T--NUDT_CHINA--collaborationsteam1.jpg|700px|center]] | ||
| + | <html> | ||
| + | </br> | ||
| + | |||
| + | <!--标题--> | ||
| + | <h2> | ||
| + | <span><span style="color:#7f1015">Mentoring TMMU-China</span></span><hr /> | ||
| + | </h2> | ||
<!--正文--> | <!--正文--> | ||
<p style="text-indent:22pt;"> | <p style="text-indent:22pt;"> | ||
| − | <span style="line-height:2;font-family:Perpetua;font-size:18px;"> | + | <span style="line-height:2;font-family:Perpetua;font-size:18px;">It was the first time for TMMU-China to take part in iGEM, so they had something unfamiliar about it. We provided a vigorous support for their project from beginning to end.</span> |
| + | </p> | ||
| + | </br> | ||
| − | + | <!--正文--> | |
| + | <p style="text-indent:22pt;"> | ||
| + | <span style="line-height:2;font-family:Perpetua;font-size:18px;">In March of 2016, we attended a meet-up hosted by The Third Military Medical University (TMMU) to discuss on some issues about iGEM.</span> | ||
| + | </p> | ||
| + | </br> | ||
| − | |||
| − | + | <!--正文--> | |
| + | <p style="text-indent:22pt;"> | ||
| + | <span style="line-height:2;font-family:Perpetua;font-size:18px;">We introduced the concepts and rules for iGEM in detail, and we also talked about some crucial points that should be paid attention to. Having had two years’ experience of taking part in the iGEM, we also shared our own lessons that we got from iGEM.</span> | ||
</p> | </p> | ||
</br> | </br> | ||
| Line 423: | Line 465: | ||
<!--正文--> | <!--正文--> | ||
<p style="text-indent:22pt;"> | <p style="text-indent:22pt;"> | ||
| − | <span style="line-height:2;font-family:Perpetua;font-size:18px;"> | + | <span style="line-height:2;font-family:Perpetua;font-size:18px;">During the meet-up, both teams communicated and exchanged each other’s ideas through presentation, brainstorm and other methods. This discussion had brought us some inspiration for our own project this year.</span> |
| + | </p> | ||
| + | </br> | ||
| − | connection between us has accelerated the project's improvement. Meanwhile, it | + | <!--正文--> |
| − | + | <p style="text-indent:22pt;"> | |
| − | let us understand one of iGEM’s core value---collaboration, sharing and | + | <span style="line-height:2;font-family:Perpetua;font-size:18px;">The connection between us has accelerated the project's improvement. Meanwhile, it let us understand one of iGEM’s core value---collaboration, sharing and community.</span> |
| − | + | ||
| − | community. </span> | + | |
</p> | </p> | ||
</br> | </br> | ||
| + | <!--插入图片--> | ||
| + | </br> | ||
| + | </html> | ||
| + | [[File:T--NUDT_CHINA--collaborationsteam2.jpg|700px|center]] | ||
| + | <html> | ||
| + | </br> | ||
| + | <!--插入图片--> | ||
| + | </br> | ||
| + | </html> | ||
| + | [[File:T--NUDT_CHINA--collaborationsfig3.jpg|700px|center]] | ||
| + | <html> | ||
| + | </br> | ||
| + | <!--插入图片--> | ||
| + | </br> | ||
| + | </html> | ||
| + | [[File:T--NUDT_CHINA--collaborationsfig4.jpg|700px|center]] | ||
| + | <html> | ||
| + | </br> | ||
| + | <!--标题--> | ||
| + | <h2> | ||
| + | <span><span style="color:#7f1015">Collaboration with NEFU_China</span></span><hr /> | ||
| + | </h2> | ||
| − | < | + | <!--正文--> |
| − | + | <p style="text-indent:22pt;"> | |
| − | + | <span style="line-height:2;font-family:Perpetua;font-size:18px;">In August of this year, we went to NEFU_China and carried out an issue communication. They took advantage of magnet tool to purify protein, which would improve the yield and purity of some protein. This idea may help us reduce the complexity of the protein cas9 purification, so as to focus more on the miRNA detection method developments. Besides, they are also very interested in our project and describe it open, close to the forefront and worth their studying. Briefly, we are pleasant to have the communication with NEFU_China.</span> | |
| − | + | </p> | |
| − | + | </br> | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| + | <!--插入图片--> | ||
| + | </br> | ||
| + | </html> | ||
| + | [[File:T--NUDT_CHINA--collaborationsteam3.jpg|700px|center]] | ||
| + | <html> | ||
| + | </br> | ||
| Line 472: | Line 535: | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| Line 504: | Line 540: | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| Line 737: | Line 761: | ||
return; | return; | ||
} | } | ||
| − | + | oTop.style.display="none"; | |
acceleration = acceleration || 0.1; | acceleration = acceleration || 0.1; | ||
time = time || 16; | time = time || 16; | ||
Revision as of 13:45, 19 October 2016
TOP