Wikiinterner Link Team:LMU-TUM_Munich/Materials (As described in the Materials section)
Wikiexterner Link Visit W3Schools
Visit W3Schools
Introduction
Purpose
Any cell mesh produced based on our bioink will suffer from exposition to serum-containing media or, in the case of in vivo application, the patient’s body fluids because of biotinidase. This ubiquitous enzyme is able to cleave biotin off of lysine – at a lower rate also off of intact protein. We may now shift this hydrolysis in two directions: either to prevent it altogether and thus stabilize our cell mesh, or to accelerate it to make our mesh quickly disintegrable. To this end, we must consider how biotin is linked to the proteins in our bioink.
NHS Esters as Amine Coupling Reagents
N-hydroxysuccinimide (NHS) esters belong to the group of amine-reactive active esters and are thus of great use in labeling proteins. Typically, NHS esters react with the primary amino group in the side chain of surficial lysine, forming a stable peptide bond.[1]
The utility of NHS esters was first suggested by Anderson et al.[2] in the context of peptide synthesis; nowadays, a plethora of NHS esters are widely used in bioconjugate techniques and thus commercially available, including those of many fluorescent dyes as well as that of biotin, often referred to as biotin-NHS (Figure 1).[3] NHS esters are typically synthesized from the corresponding carboxylic ester and NHS via the one-pot DCC coupling procedure[1] and purified by crystallization[2][3].
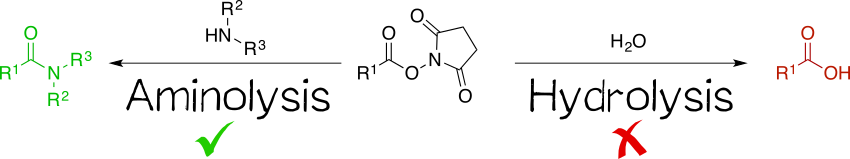
Being amine reactive, NHS esters undergo an SN2t reaction with primary or secondary amines to afford an amide bond as well as free NHS (Scheme 1, left). While this aminolysis pathway is also used in chemical synthesis, e.g. of peptides, it is key for the above-mentioned protein-labeling capability of NHS esters. The latter, however, strictly requires aqueous reaction media, so that the undesirable hydrolysis, i.e. reaction with water to give the no longer reactive carboxylic acid (Scheme 1, right), becomes a considerable side reaction even though reaction with amines, being more nucleophilic, is favored. At pH 7.0 and 0 °C, for instance, hydrolysis half-life is 4 to 5 h. For this reason, protein labeling must needs employ a great excess of NHS ester.[1][2][4][5]
For applications routinely requiring derivatization of primary amines, NHS esters are advantageous for a number of reasons. Anderson et al.[2] describe that they are highly reactive and thus give good yields in amine couplings; moreover, being crystalline, they are easy to handle. Good yields persist even when conducting the coupling reaction in the presence of water, which sets NHS esters apart from several other active esters.[4] An important benefit of NHS esters is that, in their crystalline form, they are relatively resistant to hydrolysis and thus the concomitant inactivation, meaning that they can be stored for an extended period of time under dry conditions without special precautions while retaining their activity.[3] However, this comes with a caveat: solutions of NHS esters are susceptible to hydrolysis especially if prepared in N,N-dimethylformamide or dimethyl sulfoxide. These hygroscopic solvents tend to draw moisture, promoting hydrolysis of the NHS ester; such solutions should thus be prepared immediately before use.[6]
In our work, NHS esters containing a biotin moiety, biotin-NHS among them, are used to biotinylate inanimate protein components of bioink or reservoir, for instance PAS-Lysine. For the printed mesh to be stable, this covalently attached biotin must not be cleaved. In this respect, however, we now encounter a challenge: our printing system involves live cells. These cells require cell culture media, which commonly contain serum – and serum contains the potential downfall to mesh stability: the enzyme biotinidase.
How Stable is Biotin Coupled to Proteins?
Biotinidase Hydrolytically Cleaves Biotin Off of Proteins
In the human body, biotin is a vital cofactor for several carboxylases, where it is attached to the ε-amino group of a lysine residue within the conserved sequence AMKM.[7][8] To liberate this covalently attached biotin, biotinidase, or biotin-amide amidohydrolase (EC 3.5.1.12), naturally hydrolyzes biocytin (ε-N-biotinyllysine) to biotin (vitamin H) and lysine.[7][9] The monomeric enzyme of size 76 kDa possesses a catalytic sulfhydryl group.[7][10] It not only serves to recycle biotin for renewed usage as well as to make protein-bound biotin in food bioavailable,[9] but also acts as a shuttle for biotin within the body[11]. Biotinidase deficiency thus causes multiple carboxylase deficiency, which, for one, entails serious developmental retardation because the body lacks the free biotin necessary to replenish its holocarboxylase pool; effective treatment is easily accomplished by administering daily doses of free biotin.[8]
Biotinidase is ubiquitous throughout the human body. Its specific activity is highest in serum (5.80±0.89 nmol min–1 mL–1[12]), followed by other organs including the liver, where the enzyme is produced.[8]
While biotinidase used to be considered fairly specific for biocytin,[9] evidence has accumulated that this is not the case[13]. Thus, it can also hydrolyze other biotin amide or ester conjugates, such as the methyl ester[14] or the conjugate with p-aminobenzoate, which forms the basis of a colorimetric activity assay[9][10].
Biotinidase is thought to primarily act on biocytin or small biotin-bearing oligopeptides in vivo, since its activity rapidly diminishes with increasing peptide length; it is ultimately unable to cleave biotin off of holocarboxylases.[7] However, other studies have shown that the enzyme is capable of acting on intact proteins, such as biotinylated IgG[15] and histones[16]. Since biotin participates in the chemical reaction catalyzed by carboxylases and is thus somewhat shielded, this discrepancy likely stems from different exposure to the surrounding medium.
The issue is this: With all biotin attached to lysine on both cell receptors and spacer molecules, we will invariably face slow hydrolysis if a cell mesh is exposed to serum-containing media or applied in vivo. Depending on the task at hand, we might desire our mesh to either be stable long-term or to be disintegrated more quickly. The bonding mode of the spacer’s biotin, which is introduced chemically, is key for driving this in either direction.
Biotinidase-Resistant Biotin Conjugates
If we are to prevent biotinidase from cleaving biotin off of biotinylated components in our bioprinting system and thus from disintegrating our mesh while simultaneously obviating use of special serum-free media, we must consider how to render the biotin bound to the spacer peptide non-hydrolyzable by biotinidase.
Biotinidase-resistant biotin conjugates have been extensively studied in the context of the cancer treatment "pretargeting" approach. Here, biotinylated IgG is bound to the tumor followed by sequential treatment with streptavidin and radionuclide-containing biotin, which offers the advantages of fast accretion as well as clearance combined with low tissue retention.[15][17] The general consensus is that steric hindrance at and adjacent to the peptide bond to be cleaved is paramount; for instance, p-aminobenzoate is cleaved off much faster than the m-isomer, and the o-isomer is not cleaved at all. Thus, resistance is acquired either if the amide nitrogen is tertiary, e.g. carrying an additional methyl group (as in biotinyl-sarcosine), or if the position next to this nitrogen carries a substituent.[18] Among the reported non-hydrolyzable biotin derivatives are those with the following substituents adjacent to the amide nitrogen: hydroxymethyl (serine), carboxylate (aspartate), ethyl (2-aminobutyric acid), isopropyl (valine), benzyl (phenylalanine); a methyl group (alanine), however, is not sufficient.[13][9]
Since increased steric bulk diminishes the affinity of streptavidin toward the biotin, the 2-aminobutyric acid derivative has been preferred over e.g. the valine conjugate because it binds to streptavidin more strongly and is less lipophilic. However, this deviation may not ultimately be relevant given the inherent exceptionally high avidity of streptavidin toward biotin.[13]
Inducing Faster Hydrolytic Cleavage?
To our knowledge, there have been no reports on how to improve the hydrolysis rate of biotinidase acting on intact proteins. We assume that the reason for the slowed or nil rate as opposed to biocytin is steric hindrance; thus, we must consider how to render the spacer peptide-bound biotin more accessible to the enzyme. Introducing an additional lysine between biotin and the target peptide’s lysine residue will increase the distance of the hydrolyzable amide bond to the bulk peptide and should thus facilitate hydrolysis. Synthesizing the biotinyllysine-NHS reagent is impossible without a protecting group on lysine’s α-nitrogen; to obviate protection and deprotection, aminocaproic acid, which is identical to lysine in structure except that it lacks the α-nitrogen, is desirable as the spacer moiety. While some sources cite this spacer as biotinidase resistant[9], it has been shown to be hydrolyzed in in vivo applications[19].
Design
Based on the considerations outlined above, we designed two biotinyl-amino acid NHS esters for covalent modification of target proteins:
- (+)-biotinyl threonine NHS ester, which should afford biotinidase-resistant covalent modification; and
- (+)-biotinyl aminocaproic acid NHS ester, which in
Experiments
Calibrating the NHS Ester Concentrations for Biotinylation
 Determining a viable NHS ester concentration to biotinylate the adsorbed BSA. A concentration series starting from a maximal 100 μM (BNHS and BCNHS) or 800 μM (BTNHS) was prepared in a 96-well plate. After treatment with ExtrAvidin-AP, the wells were exposed to a solution of p-nitrophenyl phosphate (0.5 mg mL–1), and substrate conversion was kinetically measured as the rate at which absorption at 405 nm increased (dA405/dt), signifying generation of p-nitrophenol. This rate correlates with the activity of AP, i.e. the amount of adsorbed ExtrAvidin-AP, in the respective well and thus with the level of biotinylation. It is apparent that the three NHS esters afford different levels of biotinylation after the allotted time, with the standard BNHS being less efficient than BCNHS but more so than BTNHS. The red horizontal line specifies dA405/dt ≈ 0.0004 s–1, around which value all three NHS esters possess a data point; the corresponding concentrations 5 μM (BNHS), 1.5 μM (BCNHS), and 200 μM (BTNHS) were used for subsequent experiments to ensure an approximately equal slope for all three derivatives.
Determining a viable NHS ester concentration to biotinylate the adsorbed BSA. A concentration series starting from a maximal 100 μM (BNHS and BCNHS) or 800 μM (BTNHS) was prepared in a 96-well plate. After treatment with ExtrAvidin-AP, the wells were exposed to a solution of p-nitrophenyl phosphate (0.5 mg mL–1), and substrate conversion was kinetically measured as the rate at which absorption at 405 nm increased (dA405/dt), signifying generation of p-nitrophenol. This rate correlates with the activity of AP, i.e. the amount of adsorbed ExtrAvidin-AP, in the respective well and thus with the level of biotinylation. It is apparent that the three NHS esters afford different levels of biotinylation after the allotted time, with the standard BNHS being less efficient than BCNHS but more so than BTNHS. The red horizontal line specifies dA405/dt ≈ 0.0004 s–1, around which value all three NHS esters possess a data point; the corresponding concentrations 5 μM (BNHS), 1.5 μM (BCNHS), and 200 μM (BTNHS) were used for subsequent experiments to ensure an approximately equal slope for all three derivatives.Since the level of biotinylation, which correlates with ExtrAvidin-AP activity, is evaluated via the kinetics of PNP being generated from PNPP, we have to ensure two aspects before starting the actual experiments: (a) the kinetic curve's slope should be within the dynamic range of the experimental setup, so that a decrease in biotinylation caused by biotinidase activity results in a maximal change in slope; and (b) it should be approximately equal for all three derivatives at time point i = 0 for better comparability. This was accomplished by assaying a concentration series of each biotinylation reagent in duplicates starting from 100 μM (BNHS and BCNHS) or 800 μM (BTNHS) and diluting 1:2 in the next set of wells, all the way down to 48.9 nM, and calculating the slope and its error for each set of two identical wells.
If we plot dA405/dt against c, we expect a saturation curve: with rising NHS ester concentration, the biotinylation level will first rapidly increase and then gradually approach a horizontal asymptote signifying full biotinylation of BSA. The semi-logarithmic plot of dA405/dt against log c transforms a saturation curve into a sigmoidal graph in which the central linear section is the target dynamic range (Figure x).[20]Error Calculations
Errors were assumed solely as the standard deviation of identical experiments; errors of other parameters (e.g. time and concentration) were neglected. For a set of identical experiments, arithmetic mean yi and standard deviation σi were calculated at each time point i, corresponding to absolute time xi, in the kinetic measurement. Now, we require a readout of the slope dA405/dt as well as its error. This task was accomplished by weighted linear regression (adapted from Ogrodnik et al.[21]): At each time point, we calculated the values si = σi–2, sx,i = xi · σi–2, sy,i = yi · σi–2, sx·x,i = xi2 · σi–2, and sxy,i = xi · yi · σi–2. From these, the sums S = i = 0n∑ si, Sx = i = 0n∑ sx,i, Sy = i = 0n∑ sy,i, Sx·x = i = 0n∑ sx·x,i, and Sxy = i = 0n∑ sxy,i were calculated over all time points i = 0 through n. With Δ = S · Sx·x – Sx · Sx, we now obtain the slope m = (S · Sxy – Sx · Sy) · Δ–1 and its error Δm = sqrt(S · Δ–1), both in units of s–1.
Results
Discussion
Syntheses: Performance and Mechanistic Considerations
Synthesis of (+)-biotin N-hydroxysuccinimide ester (BNHS, 1) was adapted from Götz et al.[22] The reaction proceeds via a straightforward carbodiimide coupling procedure between (+)-biotin as the carboxylic acid and NHS, being an N-hydroxylamine, as the hydroxyl group-bearing component (Scheme x).
The reaction involves two main steps: first, the acid’s carboxyl group conducts a nucleophilic attack on the carbodiimide carbon, resulting in an O-acylisourea intermediate after proton transfer; second, the coupling partner’s hydroxyl group in turn attacks the now activated carboxyl carbon in an SN2t reaction followed by proton transfer, yielding the desired NHS ester as well as the byproduct dicyclohexylurea, which is sparingly soluble in DMF and thus for the most part precipitates out of solution[22][23]. As such, a large part of this byproduct is easily removed by filtering the reaction mixture before removing the solvent.
DCC coupling reactions are reportedly frequently encumbered by the desired, activated intermediate O-acylisourea rearranging to the no longer reactive N-acylurea[23] (Scheme x), especially when using DMF as the solvent[24]. Nevertheless, this does not pose a problem in coupling reactions with NHS: the latter reacts with the O-acylurea so fast that rearrangement is effectively suppressed.[25] Incidentally, the Steglich esterification makes use of this same principle that adding a good nucleophile – 4-dimethylaminopyridine (DMAP) in the typical procedure – to DCC coupling reactions streamlines the reaction progress toward the desired product.[26]
Boiling the crude solid in isopropanol serves to purify the target compound, as this solvent dissolves remaining dicyclohexylurea[23] and NHS[27]. Provided that the isopropanol is dry, the NHS ester is not compromised by this procedure, being reasonably stable toward alcohols; even methanolysis – with methanol being much less sterically demanding than isopropanol – requires several days at elevated temperatures to complete.[28][29] BNHS was obtained as a white, powdery solid in 90% yield and good purity (TLC, 1H NMR). Judging by the relative peak integrals of biotin and NHS moiety, the product contains some biotin, and the slightly elevated integral of the aliphatic peaks indicates that trace isopropanol may be left from trituration.
Adapted from Bellia et al.[30] and Wilbur et al.[13], BNHS served as the starting material for synthesizing the target NHS esters of biotin derivatives with the amino acids aminocaproic acid and threonine. The mechanism of peptide bond formation proceeds via an SN2t reaction at the active ester[5] with NHS as the leaving group (Scheme 3).
While the reaction could proceed without added base triethylamine, this accelerates the reaction.[5] We must needs consider the efficiency of this coupling reaction, seeing as the amino acid is pre-dissolved in water. Under these conditions, the desired aminolysis and the non-productive hydrolysis become competing reactions, especially since both are faster in the presence of base.[5] Besselink et al. investigated the relative rates on N-Boc-protected amino acid NHS esters in a solvent mixture of DMSO and water, concluding that hydrolysis only becomes appreciable if water content exceeds 30%.[31] With a maximum of 3 mL water in 15 mL DMF, our experimental setup never surpassed 17%, so that the desired biotinyl amino acid is expected to be the major product. Coupling with aminocaproic acid is fairly straightforward, since the carboxylate group is much less nucleophilic than the amine and as such does not interfere with the reaction. Nevertheless, we should briefly consider the selectivity of coupling with threonine, which contains an additional hydroxyl group. Intuitively, we expect the amine to react preferentially, since it is more nucleophilic than the alcohol; indeed, as mentioned above, NHS esters react only slowly with alcohols.[28][29] Thus, both coupling reactions should yield the desired biotinyl amino acid as the main product.
Purification of these target compounds is easily achieved by washing the crude product with water, since they are not well soluble; as such, trace biotin derived from hydrolysis and NHS, the byproduct, are removed.[2][32] Triethylamine is removed with the solvent while heating under high vacuum. Both derivatives were afforded as white, powdery solids similar to BNHS in 80% (BCNHS) and 81% (BTNHS) yield and good purity (TLC).
Mechanistically, DCC-mediated coupling of the biotinyl amino acids with NHS proceeds in the same way as described above for synthesis of BNHS.References
- ↑ 1.0 1.1 1.2 Hermanson, G. T. (2013) Bioconjugate Techniques (3 ed.). Boston: Academic Press.
- ↑ 2.0 2.1 2.2 2.3 2.4 Anderson, G. W., Zimmerman, J. E., & Callahan, F. M. (1963). N-Hydroxysuccinimide Esters in Peptide Synthesis. Journal of the American Chemical Society, 85(19), 3039-3039.
- ↑ 3.0 3.1 3.2 Klykov, O., & Weller, M. G. (2015). Quantification of N-hydroxysuccinimide and N-hydroxysulfosuccinimide by hydrophilic interaction chromatography (HILIC). Analytical Methods, 7(15), 6443-6448.
- ↑ 4.0 4.1 Anderson, G. W., Zimmerman, J. E., & Callahan, F. M. (1964). The Use of Esters of N-Hydroxysuccinimide in Peptide Synthesis. Journal of the American Chemical Society, 86(9), 1839-1842.
- ↑ 5.0 5.1 5.2 5.3 Kalkhof, S., & Sinz, A. (2008). Chances and pitfalls of chemical cross-linking with amine-reactive N-hydroxysuccinimide esters. Analytical and Bioanalytical Chemistry, 392(1), 305-312.
- ↑ Savage, M. D. (1996). An Introduction to Avidin-Biotin Technology and Options for Biotinylation. In T. Meier & F. Fahrenholz (Eds.), A Laboratory Guide to Biotin Labeling in Biomolecule Analysis. Basel: Birkhäuser.
- ↑ 7.0 7.1 7.2 7.3 Craft, D. V., Goss, N. H., Chandramouli, N., & Wood, H. G. (1985). Purification of biotinidase from human plasma and its activity on biotinyl peptides. Biochemistry, 24(10), 2471-2476.
- ↑ 8.0 8.1 8.2 Wolf, B. (2010). Clinical issues and frequent questions about biotinidase deficiency. Molecular Genetics and Metabolism, 100(1), 6-13.
- ↑ 9.0 9.1 9.2 9.3 9.4 9.5 Wolf, B., Hymes, J., & Heard, G. S. (1990). [11] Biotinidase. In W. Meir & A. B. Edward (Eds.), Methods in Enzymology (Vol. 184, pp. 103-111): Academic Press.
- ↑ 10.0 10.1 Knappe, J., Bruemer, W., & Biederbick, K. (1963). Reinigung und Eigenschaften der Biotinidase aus Schweinenieren und Lactobacillus casei. Biochemische Zeitschrift, 338, 599-613.
- ↑ Chauhan, J., & Dakshinamurti, K. (1988). Role of human serum biotinidase as biotin-binding protein. Biochemical Journal, 256(1), 265-270.
- ↑ Wolf, B., Grier, R. E., Secor McVoy, J. R., & Heard, G. S. (1985). Biotinidase deficiency: A novel vitamin recycling defect. Journal of Inherited Metabolic Disease, 8(1), 53-58.
- ↑ 13.0 13.1 13.2 13.3 Wilbur, D. S., Hamlin, D. K., & Chyan, M.-K. (2006). Biotin Reagents for Antibody Pretargeting. 7. Investigation of Chemically Inert Biotinidase Blocking Functionalities for Synthetic Utility. Bioconjugate Chemistry, 17(6), 1514-1522.
- ↑ Pispa, J. (1965). Animal Biotinidase. Annales Medicinae Experimentalis et Biologiae Fenniae, 43, 1-39.
- ↑ 15.0 15.1 Bogusiewicz, A., Mock, N. I., & Mock, D. M. (2004). Release of biotin from biotinylated proteins occurs enzymatically and nonenzymatically in human plasma. Analytical Biochemistry, 331(2), 260-266.
- ↑ Ballard, D. T., Wolff, J., Griffin, B. J., Stanley, S. J., van Calcar, S., & Zempleni, J. (2002). Biotinidase catalyzes debiotinylation of histones. European Journal of Nutrition, 41(2), 78-84.
- ↑ Goldenberg, D. M., Chatal, J.-F., Barbet, J., Boerman, O., & Sharkey, R. M. (2007). Cancer Imaging and Therapy with Bispecific Antibody Pretargeting. Update on cancer therapeutics, 2(1), 19-31.
- ↑ Chauhan, J., & Dakshinamurti, K. (1986). Purification and characterization of human serum biotinidase. Journal of Biological Chemistry, 261(9), 4268-4275.
- ↑ Axworthy, D. B., Theodore, L. J., Gustavson, L. M., & Reno, J. M. (1999). US5955605A.
- ↑ Invitrogen. (2006). Theory of Binding Data Analysis Fluorescence Polarization Technical Resource Guide. Madison: Invitrogen Corporation.
- ↑ Ogrodnik, A., Lammers, A., Kraus, D., Fox-Beyer, B., Kämmerer, P., Smith-Gicklhorn, A., . . . Grotemeyer, J. (2007). Praktikum in Physikalischer Chemie für Studierende im Grundstudium. Munich: Chairs of Physical Chemistry at Technische Universität München.
- ↑ 22.0 22.1 Götz, M. G., James, K. E., Hansell, E., Dvořák, J., Seshaadri, A., Sojka, D., . . . Powers, J. C. (2008). Aza-peptidyl Michael Acceptors. A New Class of Potent and Selective Inhibitors of Asparaginyl Endopeptidases (Legumains) from Evolutionarily Diverse Pathogens. Journal of Medicinal Chemistry, 51(9), 2816-2832.
- ↑ 23.0 23.1 23.2 Birr, C. (1978). Aspects of the Merrifield Peptide Synthesis. Berlin: Springer-Verlag.
- ↑ Rolland, M., Amblard, M., Fehrentz, J.-A., & Martinez, J. (2002). Acylation Methods. In M. Goodman, A. Felix, L. Moroder, & C. Toniolo (Eds.), Synthesis of Peptides and Peptidomimetics (pp. 774). Stuttgart: Georg Thieme Verlag.
- ↑ Rich, D. H., & Singh, J. (1979). The Carbodiimide Method. In E. Gross & J. Meienhofer (Eds.), Major Methods of Peptide Bond Formation (pp. 241-245). New York: Academic Press.
- ↑ Neises, B., & Steglich, W. (1978). Simple Method for the Esterification of Carboxylic Acids. Angewandte Chemie International Edition in English, 17(7), 522-524.
- ↑ Knight, D. W. (2001). N-Hydroxysuccinimide. e-EROS Encyclopedia of Reagents for Organic Synthesis.
- ↑ 28.0 28.1 Gao, A., Yang, X., Zhang, C., Long, G., Pu, J., Yuan, Y., . . . Liao, F. (2012). Facile spectrophotometric assay of molar equivalents of N-hydroxysuccinimide esters of monomethoxyl poly-(ethylene glycol) derivatives. Chemistry Central Journal, 6(1), 1-9.
- ↑ 29.0 29.1 Xie, Y., Yang, X., Pu, J., Zhao, Y., Zhang, Y., Xie, G., . . . Liao, F. (2010). Homogeneous competitive assay of ligand affinities based on quenching fluorescence of tyrosine/tryptophan residues in a protein via Förster-resonance-energy-transfer. Spectrochimica Acta Part A: Molecular and Biomolecular Spectroscopy, 77(4), 869-876.
- ↑ Bellia, F., Oliveri, V., Rizzarelli, E., & Vecchio, G. (2013). New derivative of carnosine for nanoparticle assemblies. European Journal of Medicinal Chemistry, 70, 225-232.
- ↑ esselink, G. A. J., Beugeling, T., & Bantjes, A. (1993). N-Hydroxysuccinimide-activated glycine-sepharose. Applied Biochemistry and Biotechnology, 43(3), 227-246.
- ↑ Kolter, T., & Möseneder, J. (2010). Biotin ROEMPP Online. Stuttgart: Georg Thieme Verlag.