Wichtiger Kommentar: Hier wird aus Rezeptordesign hinkopiert
Design
Design of synthetic receptors presenting biotin or biotin-binding proteins
In the following, we are going to explore the thought processes that went into the design of our receptors that allow us to embed cells into a stable, three-dimensional protein matrix immediately upon printing.
For this purpose, we built two kinds of receptor constructs that both on their own, when being expressed in mammalian cells, are able to cross-link cells in streptavidin solution in our very own approach of bioprinting.
1) A receptor containing an extracellular biotin acceptor peptide that is endogenously biotinylated by a coexpressed biotin ligase (BirA) and thus presents biotin groups at the cell surface.
The biotin acceptor peptide (BAP) is a 15 amino acid peptide sequence originating from E. coli. [1] Being biotinylated by the biotin ligase BirA at a lysine residue within the recognition sequence, it mediates the functionality of the receptor by presenting biotin, allowing the interaction of the cell surface with streptavidin in the reservoir solution. The biotin ligase BirA is therefore encoded by the same vector as the receptor, with an internal ribosome entry site (IRES) allowing translation of two open reading frames (ORFs) from a single polycistronic plasmid. BirA is targeted to the ER via an Igκ signal peptide as well as an ER retention signal, allowing it to biotinylate translocating proteins possessing the recognition sequence.
2) Receptors presenting an extracellular, monomeric or single-chain avidin variant.
These avidin derivates, by design, allow the functional fusion of the otherwise tetrameric avidin molecule with our receptor. Two different variants were hereby used: The so-called enhanced monomeric avidin, a single subunit avidin that is able to bind biotin as a monomer, and single-chain avidin, which resembles the naturally occuring avidin tetramer, but has the subunits being connected via polypeptide linkers.
Both receptors generally mediate the same purpose: By interacting with streptavidin in the printing reservoir - either directly by presenting biotin (biotin acceptor-peptide) or indirectly via binding to a biotinylated linker (single chain-avidin variants), cells are being cross-linked due to the polyvalent binding of single streptavidin molecules in the printing solution to several cellular biotin groups at once. </div>
The incredible journey: signal peptides and protein targeting
Essential to a protein's function is not only its activity or binding properties, but also its presence at the right time, at the right place - in our case, the right place being the cell surface. One of the most ubiquitous elements for protein targeting is the signal peptide, which enables the translocation of proteins to the ER, from where their journey may continue to their place of function. Proteins containing a signal peptide include transmembrane proteins, secreted proteins, proteins of the ER itself, proteins of the Golgi apparatus and several more.[2] For our project, signal peptides constitute a crucial part for many constructs, being used for targeting of transmembrane proteins to the cell surface (receptors), targeting proteins to the ER (BirA) and the secretion of proteins into the surrounding medium (luciferase assays for the quantification of expression levels).
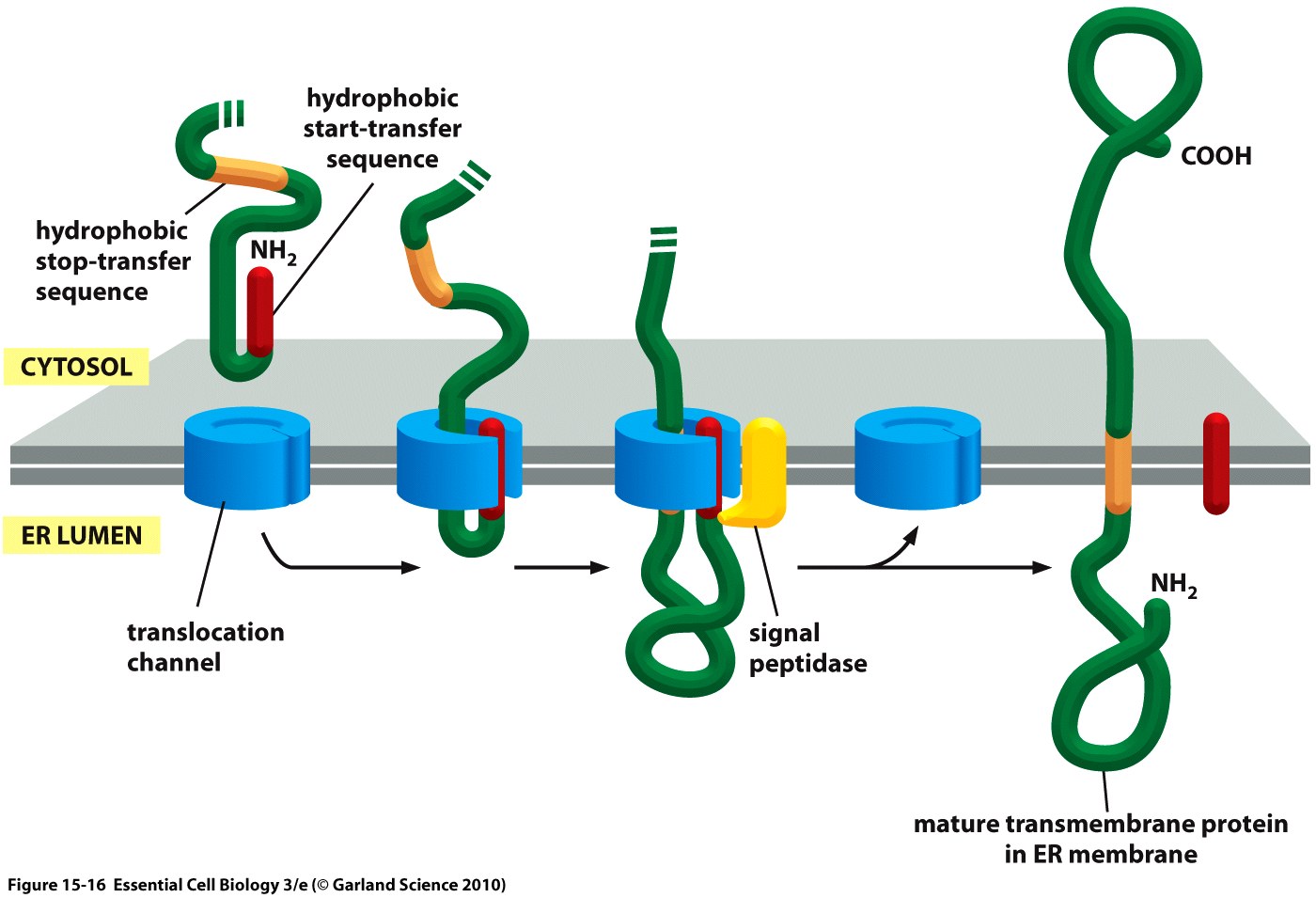
The hydrophobic signal peptide (also called start-transfer sequence) usually consists of less than 30 amino acids [4] and is recognized by the signal recognition particle upon translation at the rough ER, mediating the co-translational translocation of the nascent peptide chain into the ER.[5] During this project, three different signal peptide constructs are being tested in order to determine the one resulting in the highest expression levels:
- The EGFR signal peptide was taken from the iGEM parts registry ([http://parts.igem.org/Part:BBa_K157001:Design BBa_K157001]) and combined with a BioBrick containing the CMV promoter sequence ([http://parts.igem.org/Part:BBa_K747096 BBa_K747096]) via the RFC10 cloning standard.
- The BM40 and Igκ signal peptides were designed by us and synthesized under the aspect of adding additional spacing and functional elements to the 5'-UTR of the constructs, and combined with a BioBrick containing the CMV promoter sequence ([http://parts.igem.org/Part:BBa_K747096 BBa_K747096]) via the RFC10 cloning standard.
As shown below, the EGFR signal peptide taken from the parts registry is constructed in a way that the signal peptide ORF immediately follows the RFC10 cloning scar after the CMV promoter, thus resulting in a very short 5’ untranslated region (UTR) of the transcribed mRNA. The combination of the CMV promoter with the synthesized BM40 and Igκ signal peptides allows for a considerably longer 5’ UTR of the resulting mRNA - additionally allowing them to contain a full Kozak consensus sequence by design. The Kozak sequence is recognized by the ribosome as a translational start site; this element missing or deviating from the consensus sequence may considerably decrease translation efficiency.[6] For the EGFR signal peptide construct, a Kozak sequence is not present, as it would have to preceed the start codon ATG - a position which is occupied by the RFC10 cloning scar. Since both the distance from the promoter to the open reading frame as well as the Kozak consensus sequence are considered crucial parameters for expression levels[7], BM40 and Igκ signal peptide constructs are likely to resut in increased expression levels compared to proteins containing the EGFR signal peptide from the parts registry.
Prediction of signal peptide functionality
As the quantitative functionality of signal peptides had to be determined before including one of these into the final receptor constructs, the three options (Igκ, BM40, EGFR) were tested via bioinformatic tools as well as a secretion assay of luciferase constructs.
Bioinformatic signal peptide analysis via the SignalP server
The [http://www.cbs.dtu.dk/services/SignalP SignalP] server, being able to discriminate between the hydrophobic signal peptide sequence and the hydrophobic transmembrane domain[8], can determine whether a sequence possesses the biophysical requirements for functioning as a signal peptide. Furthermore, it is able to predict the probable cleavage site of the signal peptide after its translocation into the ER. For all constructs, signal peptide functionality is predicted, as well as a potential cleavage site. The complete translated amino acid sequences of the respective receptor constructs were used as input. The algorithm for eukaryotes with default D-cutoff value was chosen.
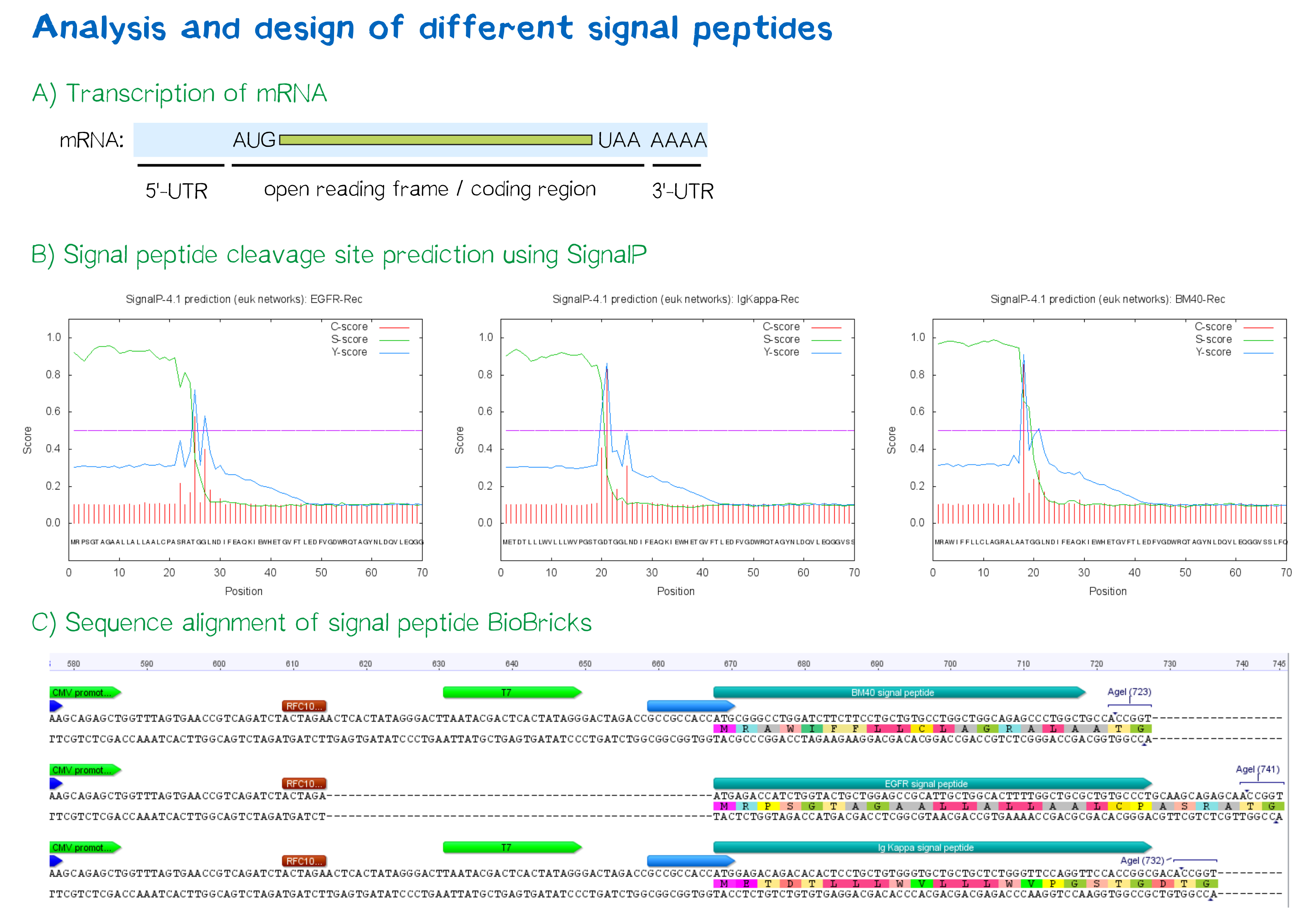
For all three signal peptides, the high S-score indicates high signal peptide functionality. For the Igκ signal peptide, a cleavage site between amino acid 20 and 21 within the signal peptide is predicted. For the BM40 signal peptide, a cleavage site between amino acid 17 and 18 is predicted, and for the EGFR signal peptide, a cleavage site between amino acid 24 and 25 is predicted.
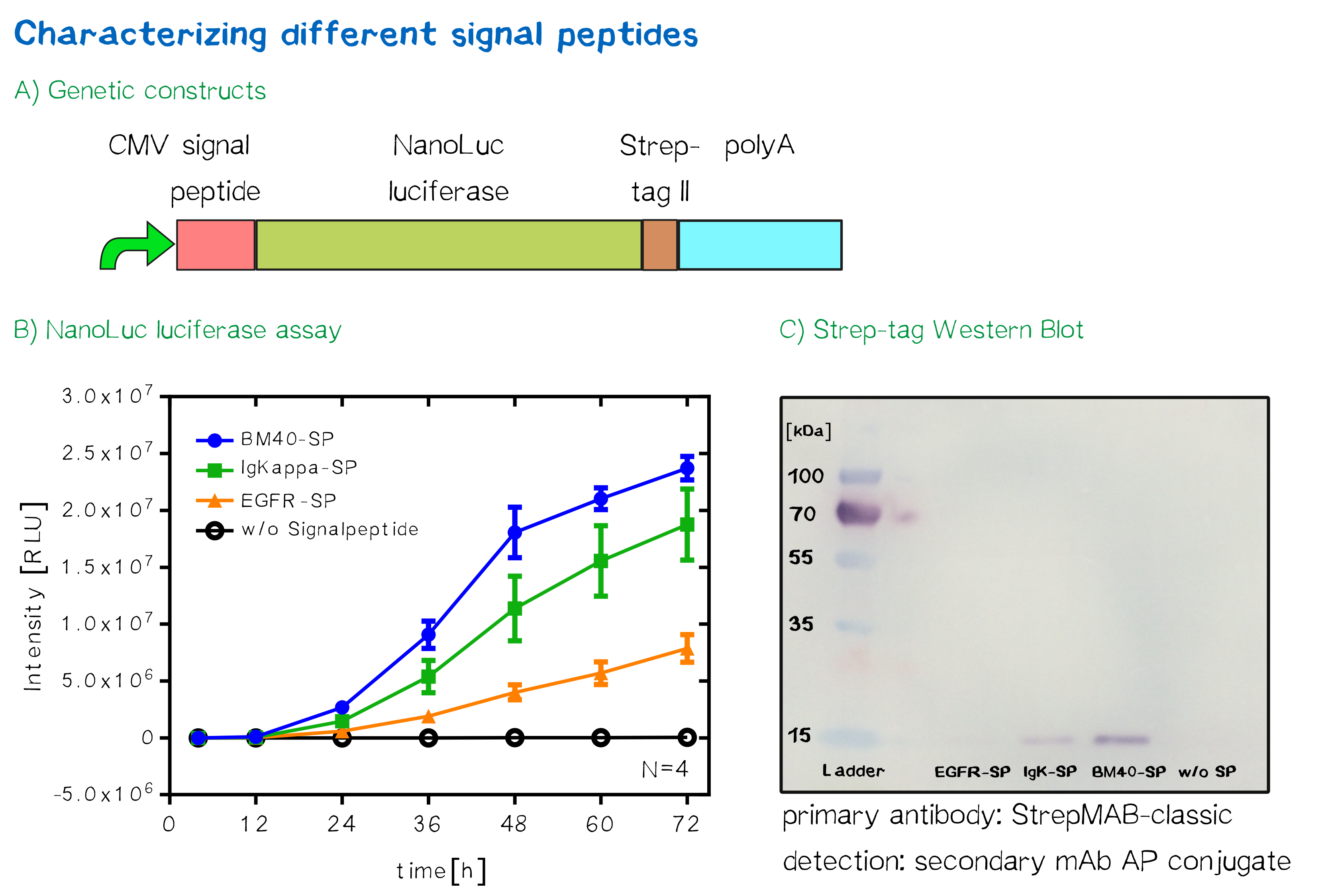
Quantification of signal-peptide mediated translocation via a secretion luciferase assay
For the quantification of signal peptide functionality via a luciferase assay, three constructs were created and tested - each containing one of three signal peptides (the EGFR signal peptide, the Igκ signal peptide or the BM40 signal peptide) and a nanoluciferase as well as a CMV promoter, a Strep-tag II for immunochemical detection and the hGH polyadenylation signal sequence. Not containing a transmembrane domain, the nanoluciferase fusion protein is being translocated into the ER and then secreted into the medium. Using a luciferase assay, one can quantifiy the amount of luminescence - and thus, proportionally, the amount of secreted luciferase - by measuring the conversion of luciferin into visible light and integrating it over a timespan of 5 s. Therefore, medium samples were taken every 12 h after transfection of cells and measured via the Promega NanoGlo luciferase assay system according to the manufacturer's instructions.
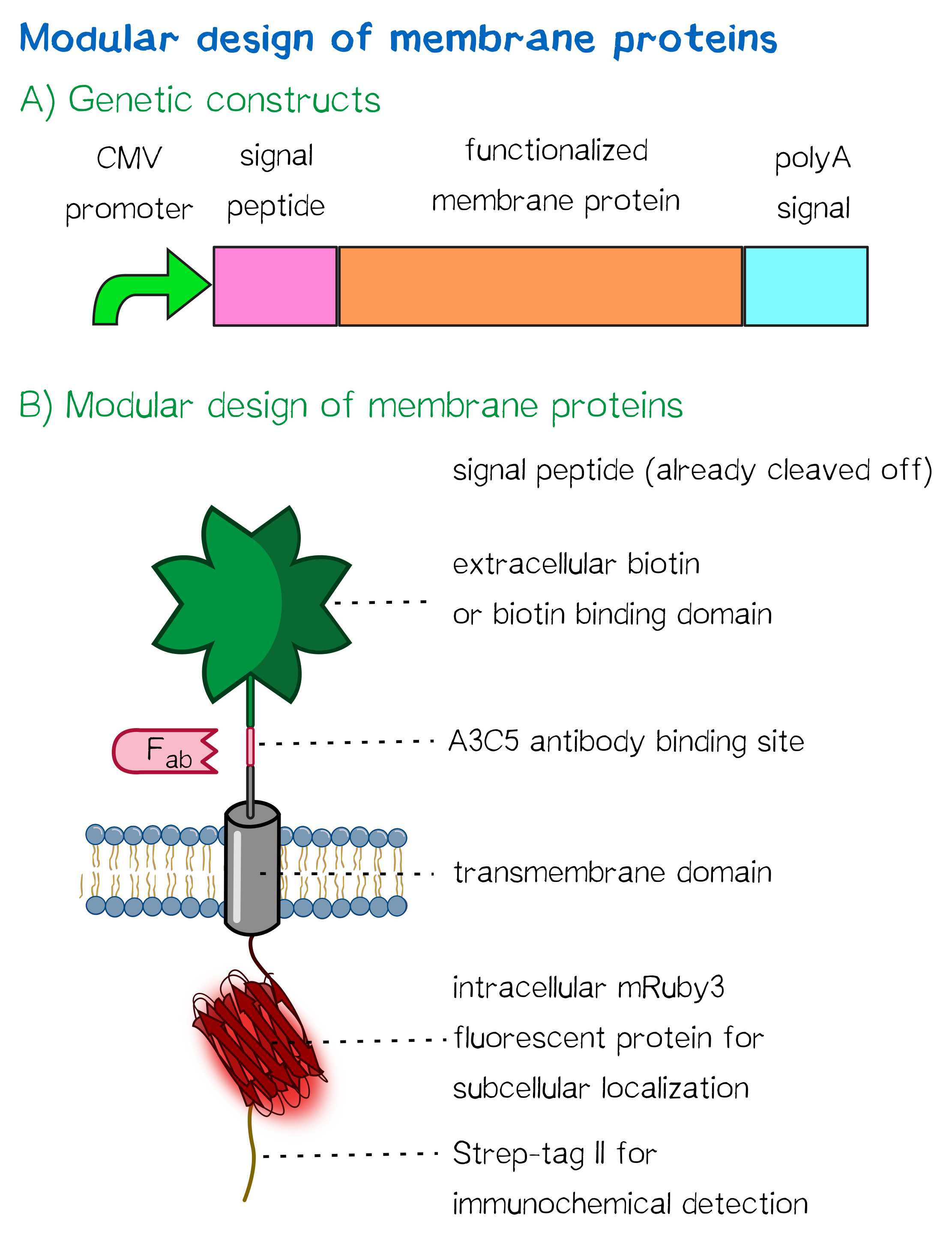
Other receptor elements: stability and detection
Apart from the functional parts that mediate the (strept)avidin-biotin interaction, other elements in the receptor were designed to make sure it is optimally translocated to the membrane, as stable as possible, and easily detectable.
EGFR transmembrane domain
For anchoring in the membrane, we decided to use a type I membrane protein, which possesses a single membrane span with defined localization of N- and C-terminus out- and inside of the cell, respectively. The N-terminal transmembrane α-helix of human EGFR (UniProt P00533, amino acids 622-653) was herefore chosen (see below the topology prediction of EGFR via the [http://www.cbs.dtu.dk/services/TMHMM/ TMHMM 2.0 server for the prediction of transmembrane helices][9]). A stop-transfer sequence consisting of charged amino acids - being characteristic for type I membrane proteins - was added at the C-terminus, and the sequence was furthermore flanked by a (GGGGC)2-linker at the N- and C-terminus, respectively. The addition of a stop-transfer sequence as in the naturally occuring EGFR sequence, as well as the addition of flexible linkers, makes our EGFR-TMD an improvement over the BioBrick [http://partsregistry.org/wiki/index.php?title=Part:BBa_K157002 BBa_K157002], which does not possess either of these elements.
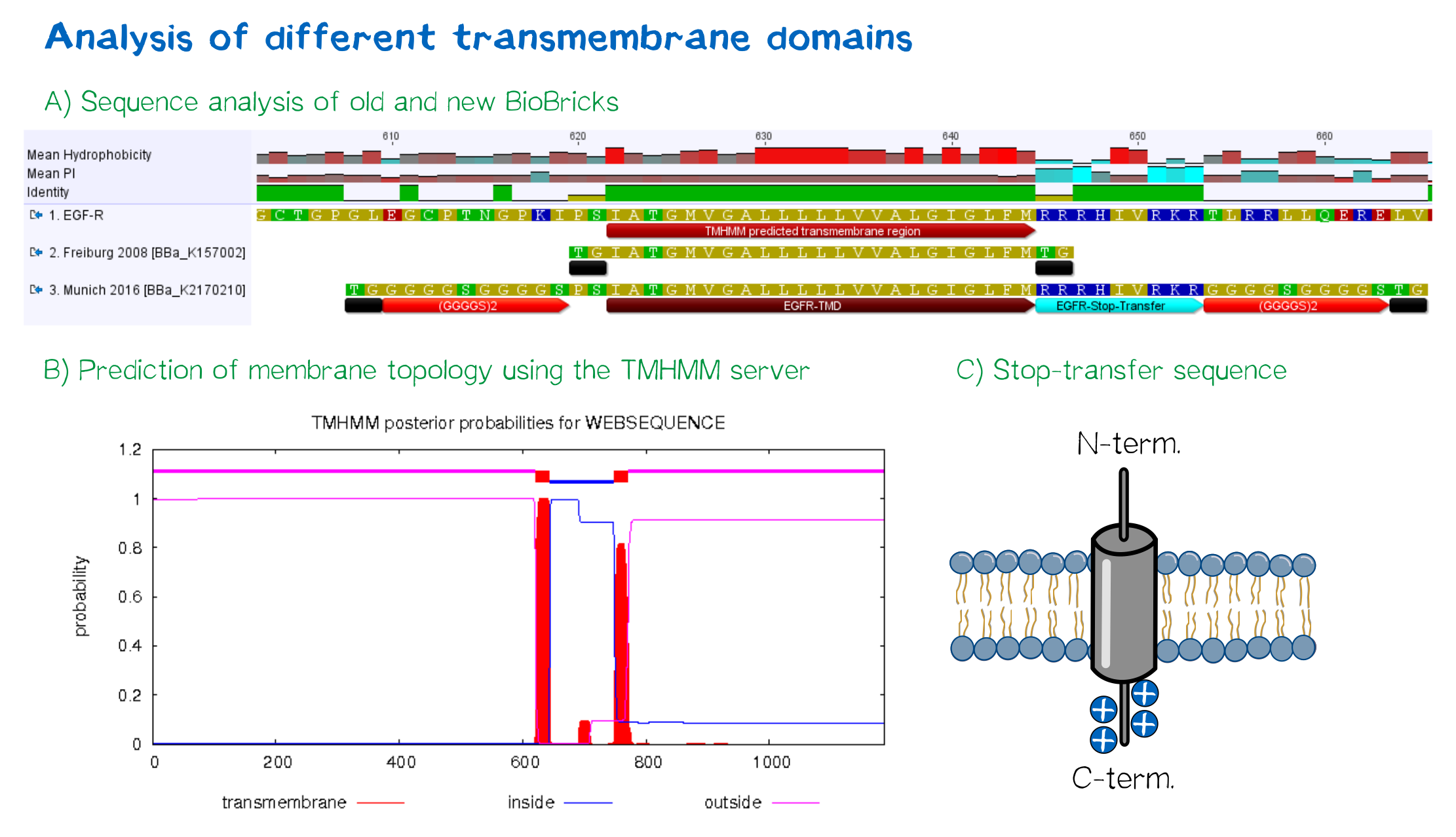
Moreover, the receptor contains three functional elements for its detection: The intracellularly located red fluorescent protein mRuby 3 for detection of the receptor via fluorescence microscopy, an extracellular epitope domain for immunochemical detection via A3C5-antibody fragments and an intracellular Strep-tag II for the detection and purification via immunochemical methods.
The vector furthermore contains the poly-adenylation signal of human growth hormone (hGH) for functional polyadenylation of the transcribed mRNA. A description for the respective functional elements is given in the following sections.
A3C5 epitope tag
Antibodies have various areas of application in life science, including protein detection, pulldown experiments and even immunotherapy. Having originally been discovered in 1995 during an in vitro screening for antibodies against cytomegaloviral proteins[10], the A3C5 antibody is an established molecular tool for specific recognition of proteins. By tagging cell surface proteins with an epitope specifically recognized by A3C5 (being a peptide sequence of 11 amino acids), one is easily able to detect tagged proteins via immunofluorescence microscopy, FACS or one may purify them via pulldown experiments. Through the latter, one may also screen for in vivo interaction partners of the tagged protein.
mRuby 3
Having originally been engineered as a monomeric form of the red fluorescent protein eqFP611, mRuby variants are sine of the brightest red fluorescent proteins available. [11] With an excitation maximum at a wavelength of 558 nm and an emission wavelength maximum of 605 nm, the resulting stokes shift of 57 nm makes mRuby a good choice for fluorescent microscopy imaging and, thus, allows the visualization of the receptor in its cellular environment by being fused to the C-terminus of the transmembrane domain. mRuby is especially powerful for the fusion with receptors - not only due to its small size and high stability, but also due to the fact that, in comparison with eGFP, it appears up to 10 times brighter in membrane enviroments. The mRuby variant used here is mRuby 3, having been engineered for even more improved brightness and photostability.[12]
Strep-tag II
The Strep-tag II is an eight amino acid peptide sequence that specifically interacts with Streptavidin and can thus be used for easy one-step purification of the receptor via immunochemical methods.[13]
Nanoluciferase
The concept of bioluminescence has a great meaning for several invertebrate species, allowing communication with other individuals, signaling receptiveness or deterring predators.[14] The most common enzyme responsible for the creation of bioluminescence are the luciferases (from the latin words ’lux’ and ’ferre’, meaning ’light-carrier’). These enzymes, among others found in fireflies and deep-sea shrimp, commonly consume a substrate called luciferin as well as energy in the form of ATP or reduction equivalents in order to create light-emission through oxidation. The underlying mechanism hereby relies on the creation of an instable, highly excited intermediate under the consumption of energy. This high-energy intermediate then spontaneously falls back into a state of lower energy, emitting the energetical difference as visible light. While mainly being used as a folded, stabilizing element at the N-terminus for the construct containing the biotin acceptor peptide, the nanoluciferase could also be used for advanced binding studies via luciferase assays.[15]

Luciferases have also found their way into biotechnological applications.[16] The simplistic concept of creating visible (and thus easily measurable) light makes luciferases ideal reporters for the expression of secretable proteins via a corresponding assay. This project also made use of a luciferase for the determination of expression levels of the BAP surface receptor while providing a stable and folded domain at the N-terminus, as well as having been used for the choice of signal peptide. The monomeric luciferase used in this project, the so-called ’NanoLuc™'[17] (due to terms of simplicity referred to as ’nanoluciferase’ in the following), can be fused to other proteins in order to make their expression visible and quantifiable. This engineered luciferase emits a steady, easily detectable glow when exposed to its substrate, while being only 19 kDa large and being brighter, more specific and steadier than other luciferases commonly found in nature. Using luciferases for the quantification of expression levels offers several advantages over fluorescent proteins, including higher sensitivity and smaller size.
CMV promoter
Last but not least, high expression levels of the receptor, so that a high avidity for the interaction may be reached. The CMV promoter (taken from the parts registry, [http://parts.igem.org/Part:BBa_K74709 BBa_K74709]) therefore enables constitutively high expression of the receptor - and, in the case of the BAP-construct, the biotin ligase BirA.

References
- ↑ Chen, I., Howarth, M., Lin, W., & Ting, A. Y. (2005). Site-specific labeling of cell surface proteins with biophysical probes using biotin ligase. Nature methods, 2(2), 99-104.
- ↑ Rapoport, T. A. (2007). Protein translocation across the eukaryotic endoplasmic reticulum and bacterial plasma membranes. Nature, 450(7170), 663-669.
- ↑ Taken from Alberts, B., Bray, D., Hopkin, K., Johnson, A., Lewis, J., Raff, M., ... & Walter, P. (2010). Essential cell biology. Garland Science.
- ↑ Walter, P., Ibrahimi, I., & Blobel, G. U. N. T. E. R. (1981). Translocation of proteins across the endoplasmic reticulum. I. Signal recognition protein (SRP) binds to in-vitro-assembled polysomes synthesizing secretory protein. The Journal of Cell Biology, 91(2), 545-550.
- ↑ Keenan, Robert J., et al. "The signal recognition particle." Annual review of biochemistry 70.1 (2001): 755-775.
- ↑ Kozak, M. (1989). The scanning model for translation: an update. The Journal of cell biology, 108(2), 229-241.
- ↑ Kozak, M. (1987). An analysis of 5'-noncoding sequences from 699 vertebrate messenger RNAs. Nucleic acids research, 15(20), 8125-8148.
- ↑ Petersen, T. N., Brunak, S., von Heijne, G., & Nielsen, H. (2011). SignalP 4.0: discriminating signal peptides from transmembrane regions. Nature methods, 8(10), 785-786.
- ↑ Sonnhammer, E. L., Von Heijne, G., & Krogh, A. (1998, July). A hidden Markov model for predicting transmembrane helices in protein sequences. In Ismb (Vol. 6, pp. 175-182).
- ↑ Alexander, H., Harpprecht, J., Podzuweit, H. G., Rautenberg, P., & Müller-Ruchholtz, W. (1994). Human monoclonal antibodies recognize early and late viral proteins of human cytomegalovirus. Human Antibodies, 5(1-2), 81-90.
- ↑ Kredel, S., Oswald, F., Nienhaus, K., Deuschle, K., Röcker, C., Wolff, M., ... & Wiedenmann, J. (2009). mRuby, a bright monomeric red fluorescent protein for labeling of subcellular structures. PloS one, 4(2), e4391.
- ↑ Bajar, B. T., Wang, E. S., Lam, A. J., Kim, B. B., Jacobs, C. L., Howe, E. S., ... & Chu, J. (2016). Improving brightness and photostability of green and red fluorescent proteins for live cell imaging and FRET reporting. Scientific reports, 6.
- ↑ Schmidt, T. G., & Skerra, A. (2007). The Strep-tag system for one-step purification and high-affinity detection or capturing of proteins. Nature protocols, 2(6), 1528-1535.
- ↑ Case, J. F. (2004). Flight studies on photic communication by the firefly Photinus pyralis. Integrative and comparative biology, 44(3), 250-258.
- ↑ Zhang, L., Song, G., Xu, T., Wu, Q. P., Shao, X. X., Liu, Y. L., ... & Guo, Z. Y. (2013). A novel ultrasensitive bioluminescent receptor-binding assay of INSL3 through chemical conjugation with nanoluciferase. Biochimie, 95(12), 2454-2459.
- ↑ Gould, S. J., & Subramani, S. (1988). Firefly luciferase as a tool in molecular and cell biology. Analytical biochemistry, 175(1), 5-13.
- ↑ Hall, M. P., Unch, J., Binkowski, B. F., Valley, M. P., Butler, B. L., Wood, M. G., ... & Robers, M. B. (2012). Engineered luciferase reporter from a deep sea shrimp utilizing a novel imidazopyrazinone substrate. ACS chemical biology, 7(11), 1848-1857.