Interaction of biotin and streptavidin leads to a pseudopolymerization of cells
For cross-linking cells upon printing using our biotINK-approach, we utilize the extraordinary strength of the biotin-streptavidin interaction. The cells express a receptor which allows them to present a biotin molecule or a streptavidin variant on their cell surface (see Receptor).
The tetrameric streptavidin protein in the printing solution is able to cross-link the biotinylated cells and thus make them form a stable network upon printing. In order to adjust intercellular distances, a biotinylated linker peptide containing regularly spaced biotinylated lysine residues is used.
Streptavidin and Biotin: The strongest non-covalent affinity in nature
Streptavidin is a tetrameric protein with a molecular weight of 52.8 kDa[1] and was originally isolated from Streptomyces avidinii. Each subunit is able to bind a single biotin molecule. This specific, non-covalent bond with a unique femtomolar dissociation constant (KD = 10-15 M-1) is one of the strongest known biological interactions.[2] Antibodies, in comparison, have dissociation constants in the range of 10-7 – 10-11 M-1 - being less affine by a factor of a million.
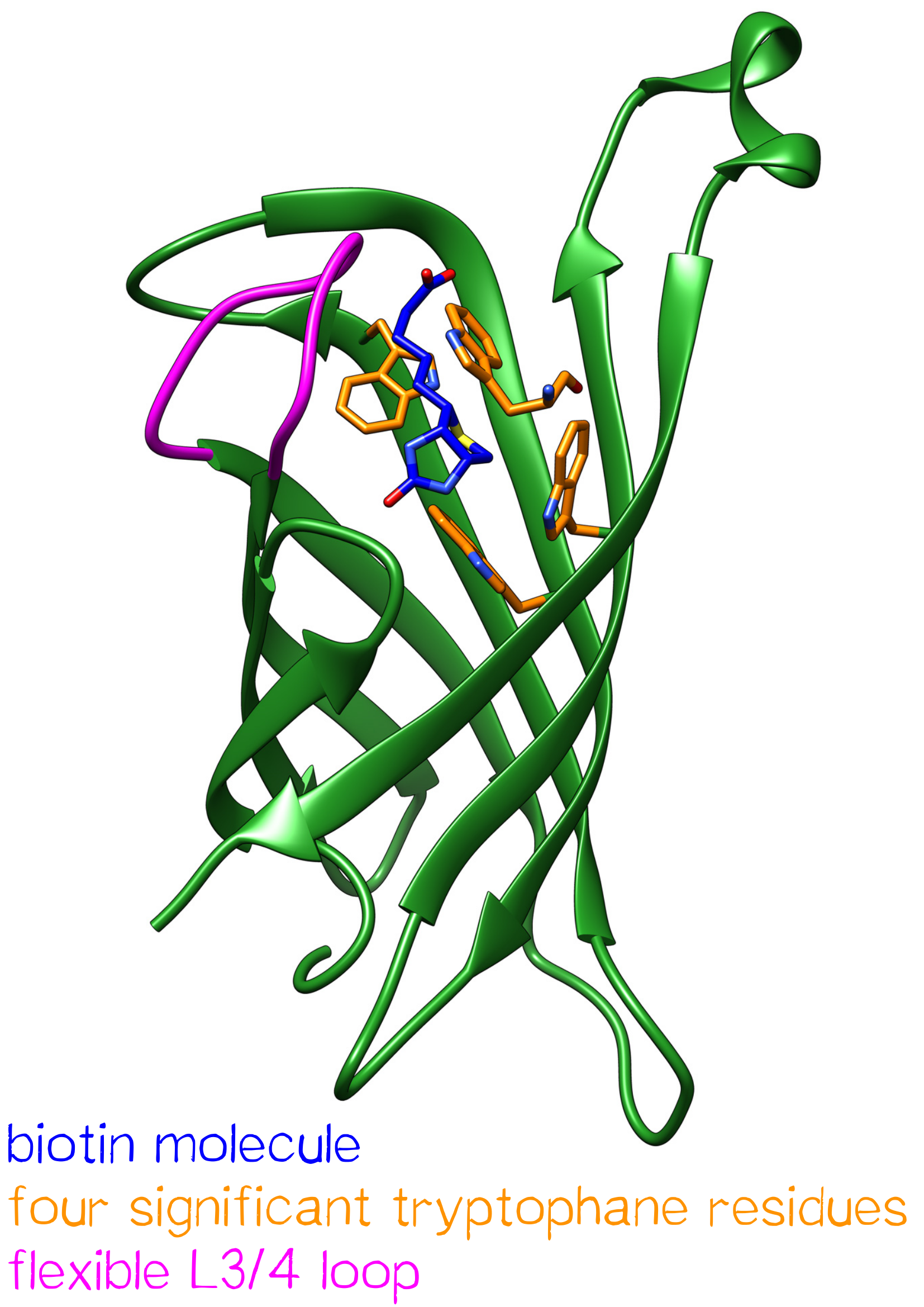
One subunit of streptavidin is organized as an eight stranded antiparallel beta sheet of coiled polypeptide chains which form a hydrogen bonded barrel with extended hydrogen loops[3] (figure 2).
The biotin binding pocket primarily consists of aromatic and polar side chains, which interact with the hetero atoms of biotin.
Due to the high specificity and affinity of the interaction, streptavidin is often used for protein-purification via affinity chromatography and as a detection and coupling reagent in daily lab routine.[4]
The rapid and very strong linkage between the two binding partners results from the creation of multiple hydrogen bonds as well as Van der Waals interactions upon binding. Comparing the Apo-state of streptavidin and the streptavidin-biotin complex hereby shows several structural differences of the binding pocket.
The ordering of a flexible surface polypeptide loop results in burying the biotin molecule in its pocket. Without biotin, the L3/4 loop is disordered and does not give clear electron density, while it closes upon biotin binding.[5]
Important for the high energetical barrier required for dissociation are biotin-tryptophan contacts in the binding pocket. There are four significant tryptophan residues involved in the binding site. Three of those (Trp79, Trp92 and Trp108) are lined up in one section. Trp120 binds the biotin molecule from an adjacent subunit and displays a much larger influence in binding free energy.[6]
Moreover the hydrogen bonding network and Van der Waals forces show great influence in binding free energy. Compared to similar hydrogen bonding donors and acceptors of other protein-ligand systems the hydrogen bonding of streptavidin to the ureido oxygen of biotin is remarkably high.
Streptavidin variants and homologs
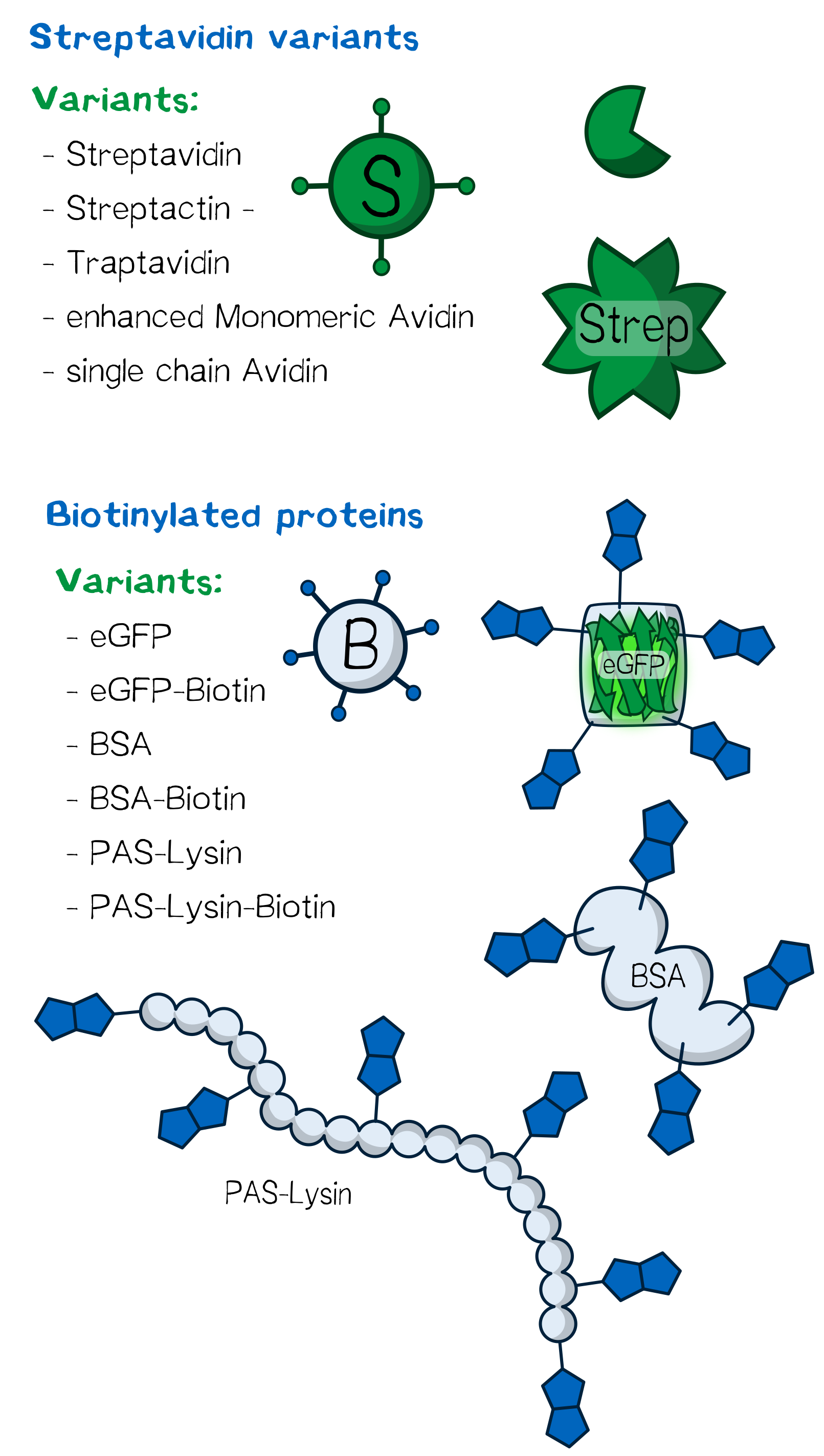
Several streptavidin variants were expressed, purified and tested in order to determine how their specific properties influence the printing processes. An overview of the different physical properties of the Streptavidin variants we used are shown below.
| protein | amino acids (without Met) | molecular weight monomer [Da] | molecular weight tetramer [Da] | theoretical isoelectric proint (pI) | extinction coefficient (monomer, ε280) [M-1 cm-1] | Tm [K] |
|---|---|---|---|---|---|---|
| Streptavidin wt | 126 | 13349.55 | 52801.36 | 6.09 | 41940 | 361.78 |
| Traptavidin | 126 | 13278.43 | 52516.88 | 5.14 | 41940 | 360.59 |
| Streptactin | 126 | 13390.65 | 52516.88 | 8.32 | 41940 | 350.86 |
| enhanced monomeric Avidin | 138 | 15369.51 | / | 5.91 | 35075 | 320.6 |
Traptavidin: increased mechanical strength of biotin binding
Being a streptavidin derivate with two point mutations that increase its affinity (S52G, R53D), traptavidin shows a 10-times lower dissociation constant compared to the wildtype (KD=~10-16 M). Moreover, the tetramer has an improved thermostability and thus remains more functional at higher temperatures (TM~10 °C higher)[7]. In contrast to streptavidin, the Traptavidin L3/4 loop (residues 45-50) does not change its conformation upon binding biotin. The binding pocket is already closed and lacks flexibility. The loss of a structural change is thought to lower the entropic cost of binding and therefore decreases the koff-value as well as the dissociation constant.
Strep-Tactin: a Strep-tag binding protein
The Strep-tag is a small peptide composed of eight amino acids and can be used as affinity tag by genetic fusion to the N- or C-terminus of a recombinant protein. Using the Strep-tag's interaction with "Strep"-Tactin for affinity chromatography enables a simple and rapid one step-purification and provides great yields in protein purification. The strep-tagged protein binds Strep-Tactin, which is immobilized on the column matrix. After removing host proteins by washing, the target protein can be eluted with desthiobiotin, which binds the Strep-Tactin several orders of magnitude stronger. Over the years both, the Strep-tag and the streptavidin were optimized resulting in Strep-tag II (amino acid sequence: WSHPQFEK) affinity tag with enhanced binding to Strep-Tactin. [8]
Enhanced Monomeric Avidin (eMA: engineered for applications like molecular labelling
Due to the monovalent nature of eMA, unwanted cross-linking of biotin conjugats can not occur.
Since streptavidin binding pockets depend on one residue from a adjacent subunit, binding affinity of a single subunit is decreased (KD = 10-7 to 10-9 M-1). eMA consists of a monomerized Rhizavidin dimer, which contains a disulfid bond in the binding site restraining the protein and forming a rigid binding pocket[9].
Production and purification of streptavidin
The minimal streptavidin wildtype gene is cloned on a pSA plasmid. The transcription is under control of a T7-promoter. The E.coli BL-R (DE3) strain codes the T7-polymerase under the control of a lacUV5 promotor, which can be induced by Isopropyl β-D-1-thiogalactopyranoside (IPTG).
A shaking flask with 2 L LB medium supplemented with ampicillin is inoculated 1:40 with a 50 ml preculture grown at 30 °C over night.
The bacterial culture shakes at 37 °C for at least four hours until its optical density reaches 0.6 and protein production is induced with 1 mM IPTG. After 4 hours the cells can be harvested by centrifugation (5000 rpm for 30 min).
The cell pellet is resuspended in a 20 mM Tris/HCl buffer (500 mM NaCl, pH 8) and Homogenization can be proceeded with French press or PANDA. The cell lysate is centrifuged at 11500 rpm for 30 min to separate the inclusion bodies in the pellet from the soluble proteins of the supernatant.
The inclusion bodies consist of aggregated and misfolded protein, those are denaturated with 6 M guanidinium chloride and centrifuged at 15000 rpm for 15 minutes.
The aggregations in the pellet are discarded and the supernatant is added slowly into a high volume of PBS (30 ml PBS for 1 ml protein solution) and incubated over night. Guanidinium chloride is diluted and proteins are able to fold correctly. The solution is centrifuged again (11500 rpm for 30 min) to remove aggregates.
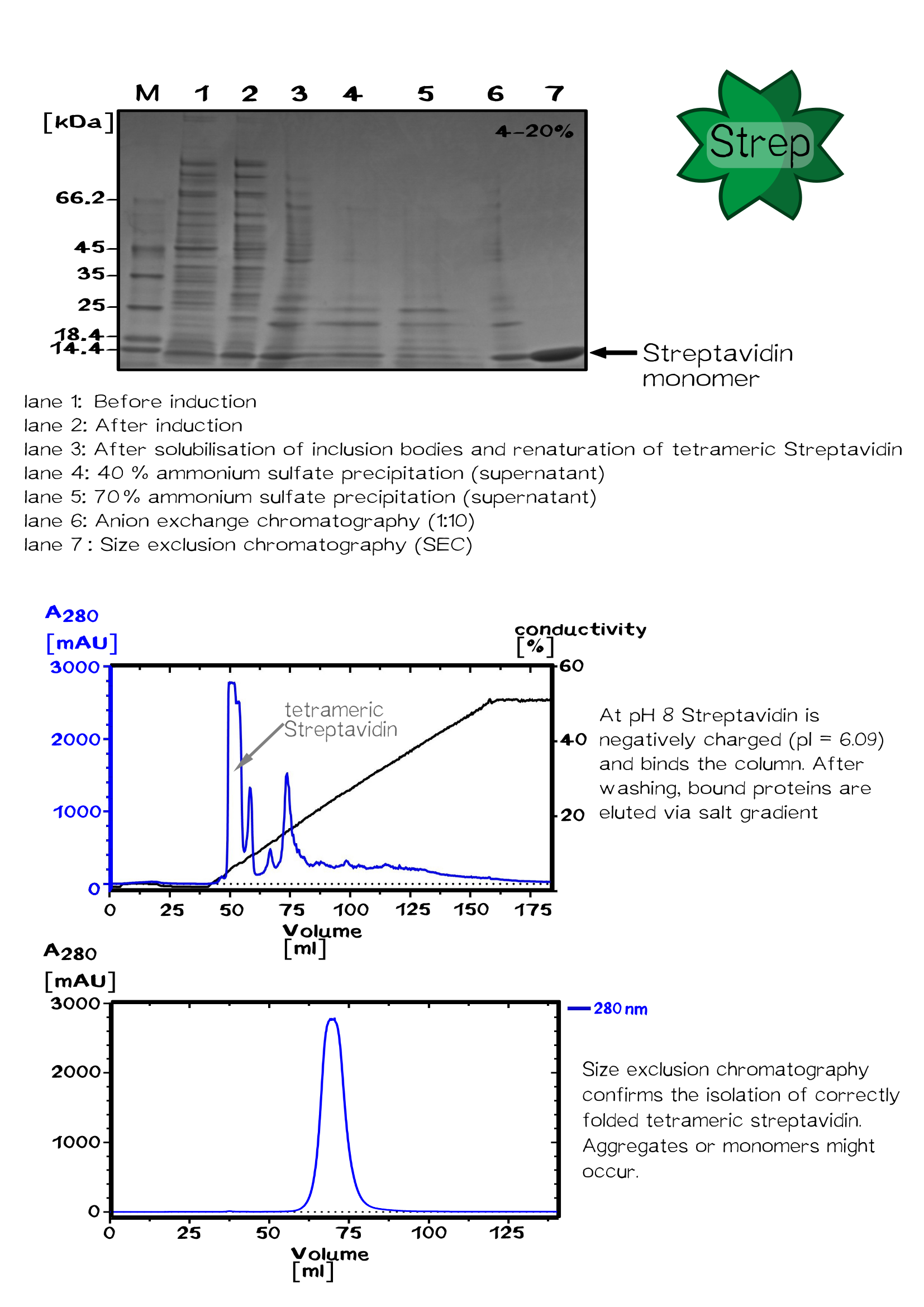
The streptavidin purification is proceeded by fractionally ammonium sulfate precipitation. After each precipitation step the solution is incubated without stirring for few hours and centrifuged at 11500 rpm for 30 minutes afterwards.
First precipitation step is 40 % saturation (1.75 M). streptavidin remains soluble, the pellet can be discarded. After centrifugation the ammonium sulfate saturation is increased up to 70 % (3.37 M), streptavidin is no longer soluble and precipitates.
The pellet is resuspended in a 50 % saturated ammonium sulfate solution (2.25 M), streptavidin again precipitates. After centrifugation the pellet is resuspended in 1x PBS buffer.
As a result the solution contains only those proteins, which precipitate between 40 % and 50 % ammonium sulfate saturation. One last centrifugation at 15000 rpm for 30 minutes removes insoluble impurities.
The solution is dialysed against 20 mM Tris/HCl buffer without salt (pH 8). Streptavidin has an isoelectric point of 6.09, therefore it is negatively charged in its buffer and a positively charged ResQ column is used for ion exchange chromatography. An increasing salt concentration of the buffer causes fractional elution of protein.
Streptavidin monomers, tetramers and aggregates can be identified by gel filtration with a sephacryl column.
After sterile filtration, a UV/VIS spectrum of the solution is measured in order to determine its concentration.[10]
The production of Traptavidin and Strep-Tactin is analogue. Enhanced monomeric Avidin is isolated via IMAC.
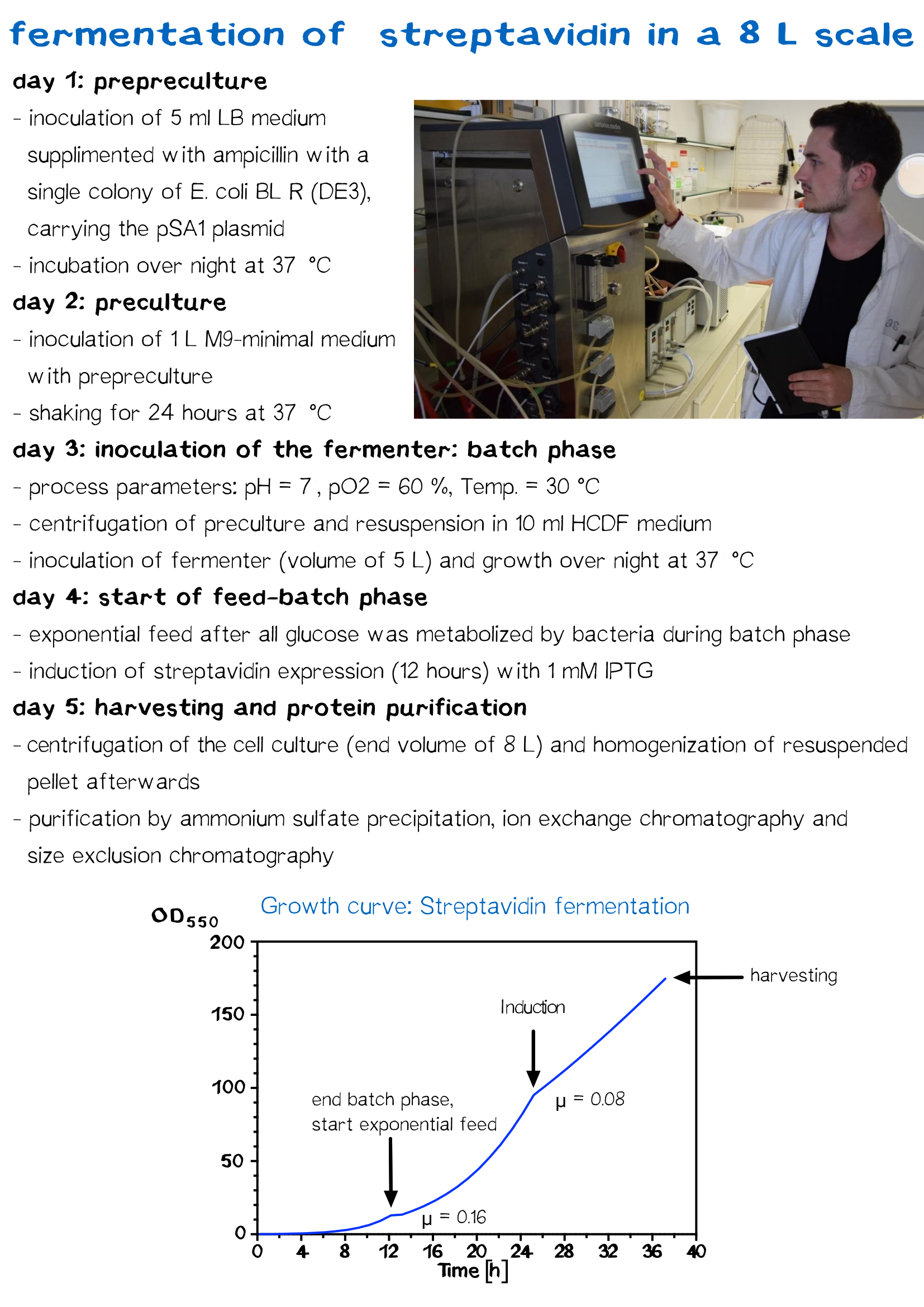
Streptavidin Fermentation
Fermentation is utilized for upscaling protein production under strictly defined conditions. Oxygen saturation of 60 %, pH 7 and a temperature of 30 °C remain constant during the whole process. Using a high cell density fidellity (HCDF) medium and applying an exponential feed results in extraordinary high cell density (OD > 60), which can not be reached in shaking flask production (OD in a range of 4).
Biotinylated PAS-lysine for cross linking
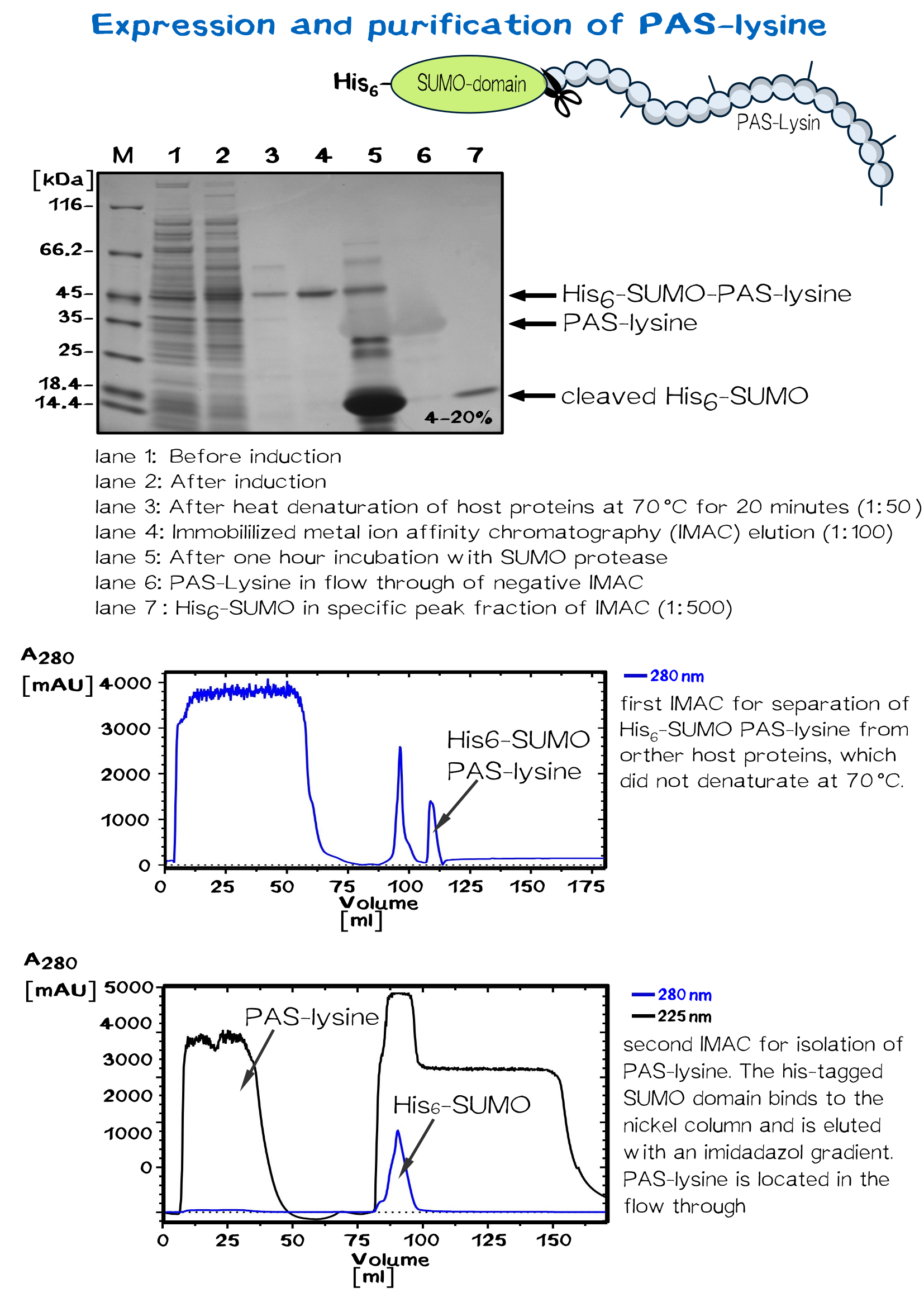
Biotinylated SUMO-PAS-lysine is used as a linker molecule in the biotink technology. The PAS-lysine linker is fused to SUMO protein domain, which carries a N-terminal His-tag and therefore provides simple purification.
PAS consists of conformationally disordered polypeptide chains with expanded hydrodynamic volume due to the small residues Pro, Ala and Ser (PAS). [11] The uncharged polymer is very good soluble and includes ten lysine residues, which allow chemically biotinylation at the terminal amino groups.
A 50 ml preculture with E.coli BL R (DE3) carrying the pSUMO plasmid is poured into a shaking flask with 2 L LB medium. After the bacterial suspension reaches an optical density of 0.5 the production of PAS-lysine is induced by adding IPTG (1 mM).
The cells are harvested (5000 rpm, 30 minutes) after five hours shaking at 37 °C and resuspended in 20 mM Tris/HCl buffer with 500 mM salt, pH 8. Homogenization is conducted by French press or PANDA. SUMO PAS-lysine is very good soluble, after centrifugation at 11500 rpm for 30 minutes it is located in the supernatant, the pellet can be discarded.
SUMO PAS-lysine has the property to remain native even at high temperatures, where most protein denaturate. Heating the solution up to 70 °C for 20 minutes and centrifugation afterwards removes nearly all other proteins.
The solution is dialysed against IMAC buffer (50 mM sodium dihydrogen phosphate and 500 mM salt, ph 7.5) and loaded on a chromatography column packed with nickel. After washing away impurities, the addition of imidazole results in elution of the SUMO PAS-lysine.
The SUMO domain has no further use and is cleaved off by the SUMO-protease ULP. Another IMAC chromatography separates the PAS-lysine from SUMO domain, which binds specifically to the column. PAS lysine can be identified in the flow through via UV absorption. Because PAS-lysine does not contain aromatic residues, its absorption at 280 nm is very low, while the absorption at 225 nm (amid bondage) is average.
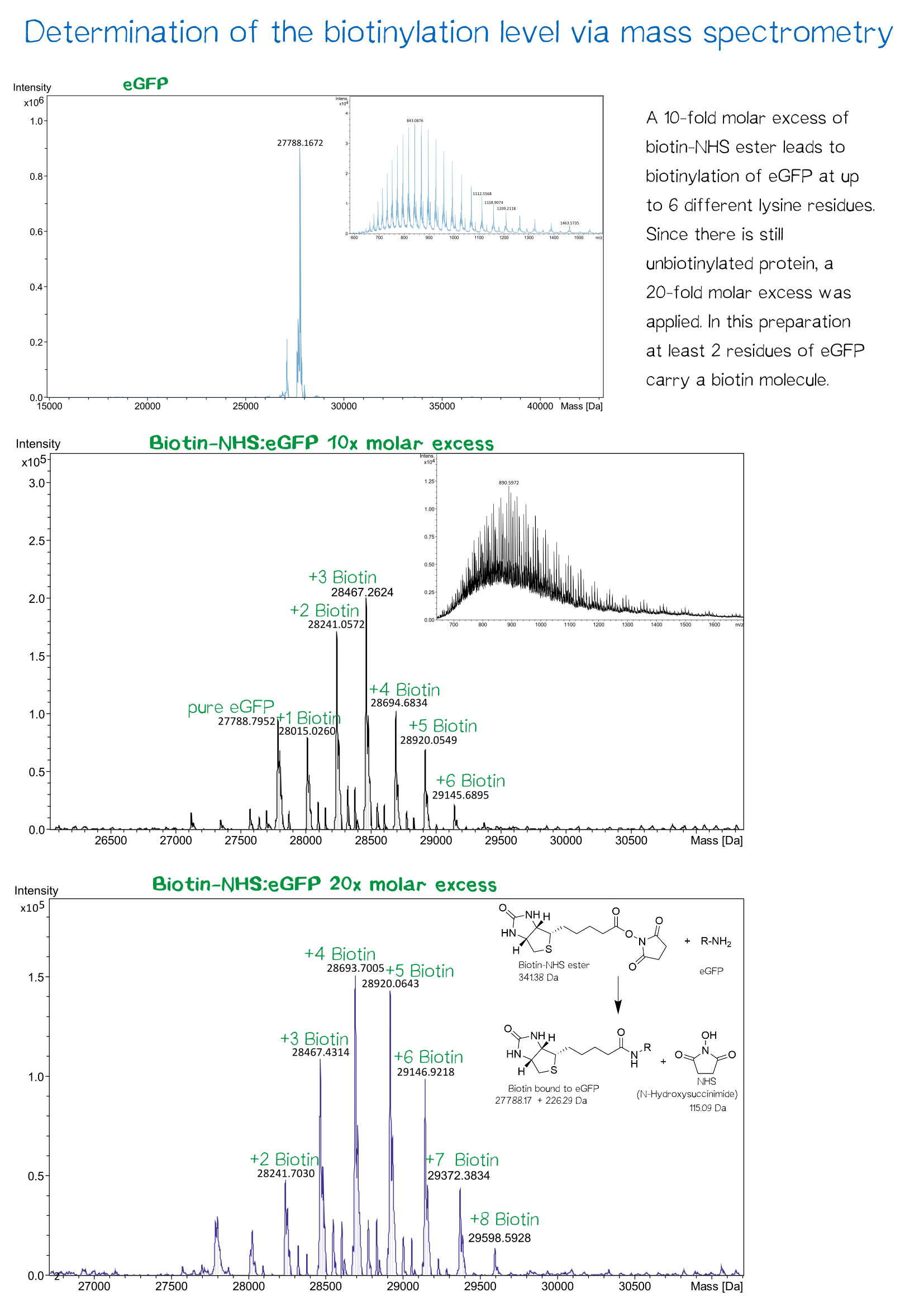
The proper fractions are pooled and the solution is dialysed against 20 mM Tris/HCl buffer with 500 mM salt, pH 8. The linker molecule can now be biotinylated in a molar excess of 10 or 20. After incubation over night, dialysis removes biotin molecules which did not bind the PAS-lysine linker.
eGFP and BSA: alternative linker molecules
Serum albumin is the most abundant plasma protein in mammals. It is a transporter molecule for a diverse range of metabolites, drugs, nutrients, metals and other molecules and shows remarkable ligand binding capacity. [12] Bovine serum albumin, originally isolated from a cow, is often used as a concentration standard for protein assays, it is therefore in stock of many laboratories. Since it has 60 lysine residues within its 607 amino acids long sequence, the soluble BSA protein works well as a linker molecule.
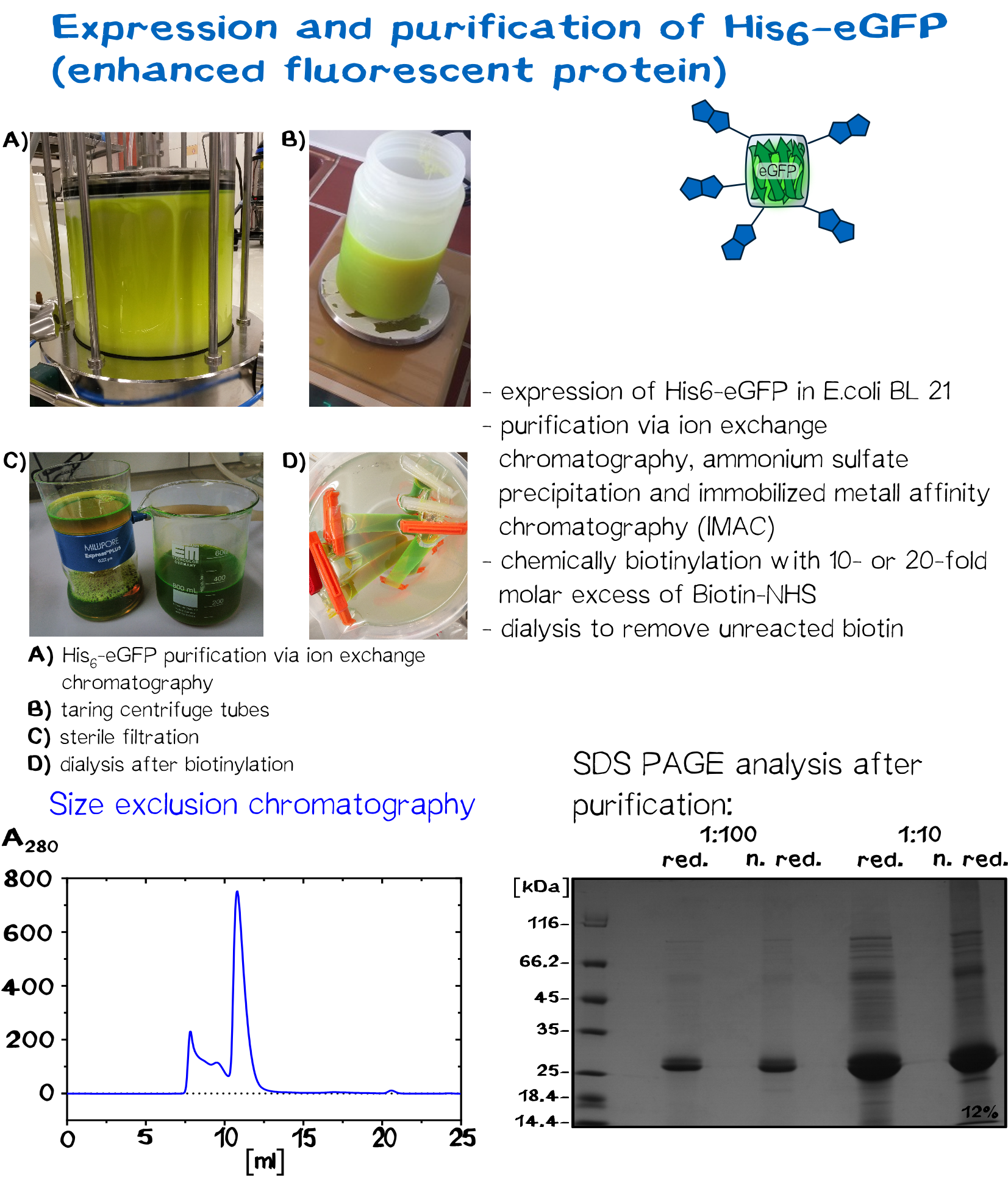
In recent years eGFP has emerged as a powerful reporter molecule for monitoring gene expression, protein localization and protein-protein interaction. [13]. Due to its emission of green light, eGFP helped improving the printing and polymerization process by enabling the opportunity to see if polymerization of the sample was successful immediately. Thus, only promissing samples were analysed under a microscope.
As a globular protein with 20 lysine residues in its sequence, eGFP can be used as a chemically biotinylated linker molecule.
Further characterisation of streptavidin variants
Overview over produced proteins
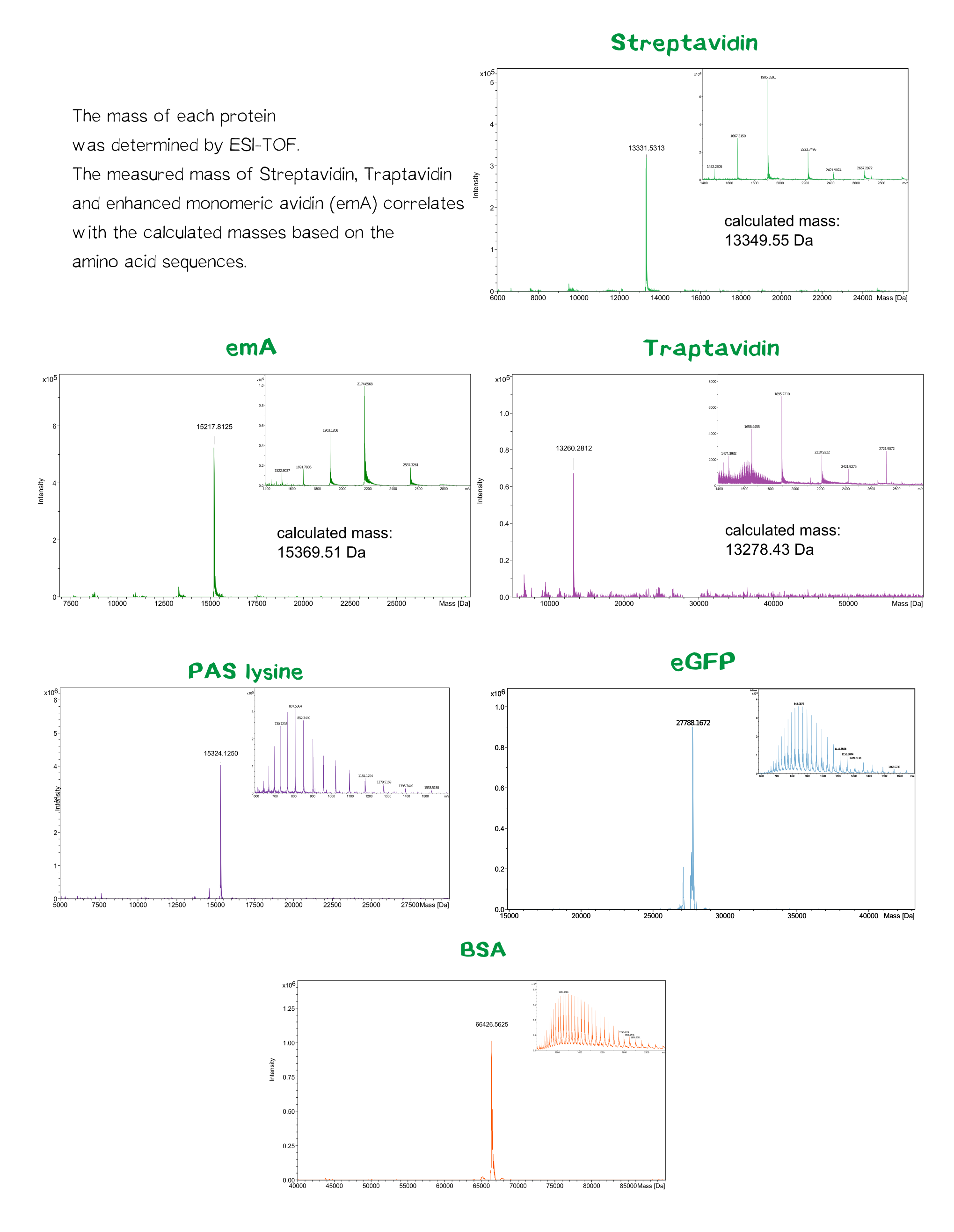
Streptavidin and its variants were produced by cytoplasmic expression in E. coli in inclusion bodies. After stabilization and refolding of the functional tetrameric and monomeric variants, purification was carried out by ammonium sulfate precipitation, Ion Exchange Chromatography and size exclusion chromatography. Samples of each required step were analysed by SDS-PAGE. Finally each of the successfully purified variants were characterized. Mass spectrometry was used to confirm the expected size, circular dichroism spectroscopy (cd) to determine folding states, fluorescence titration and surface plasmon resonance for determining binding affinities.
Results can be seen in Table 1.
Quality control using ESI-TOF mass spectrometry
Quality control using SDS-PAGE and isoelectric focusing
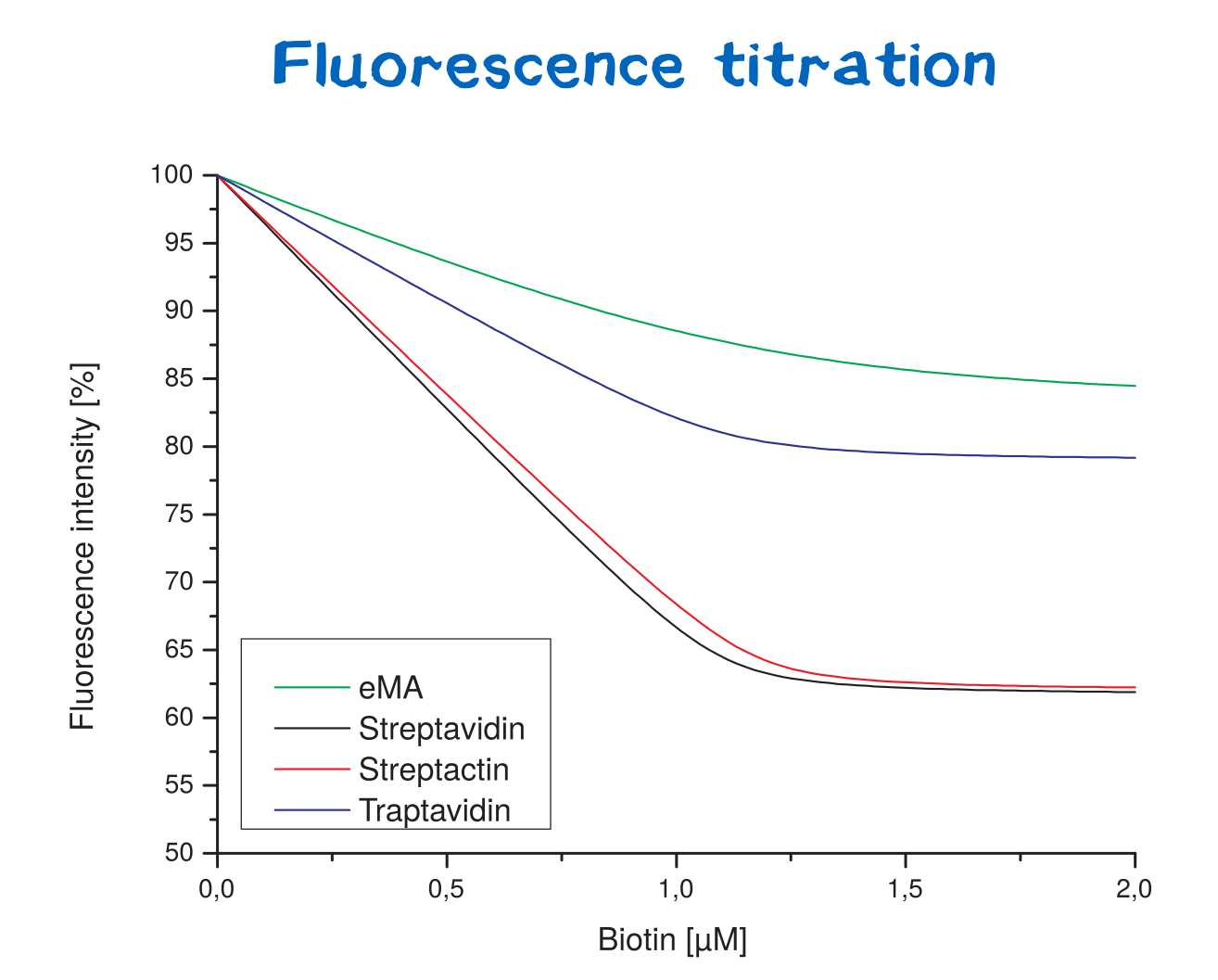
Affinity determination using fluorescence titration
As you can see in figure 13 Streptavidin and Traptavidin have a lower KD as Streptactin and emA. Unfortunately it was not possible to determine the correct KD value for each protein. Because therefore much lower concentrations than used were necessary, which the given equipment was not able to detect and measure properly. Thats why only a qulaitative evaluation of the curves were possible.
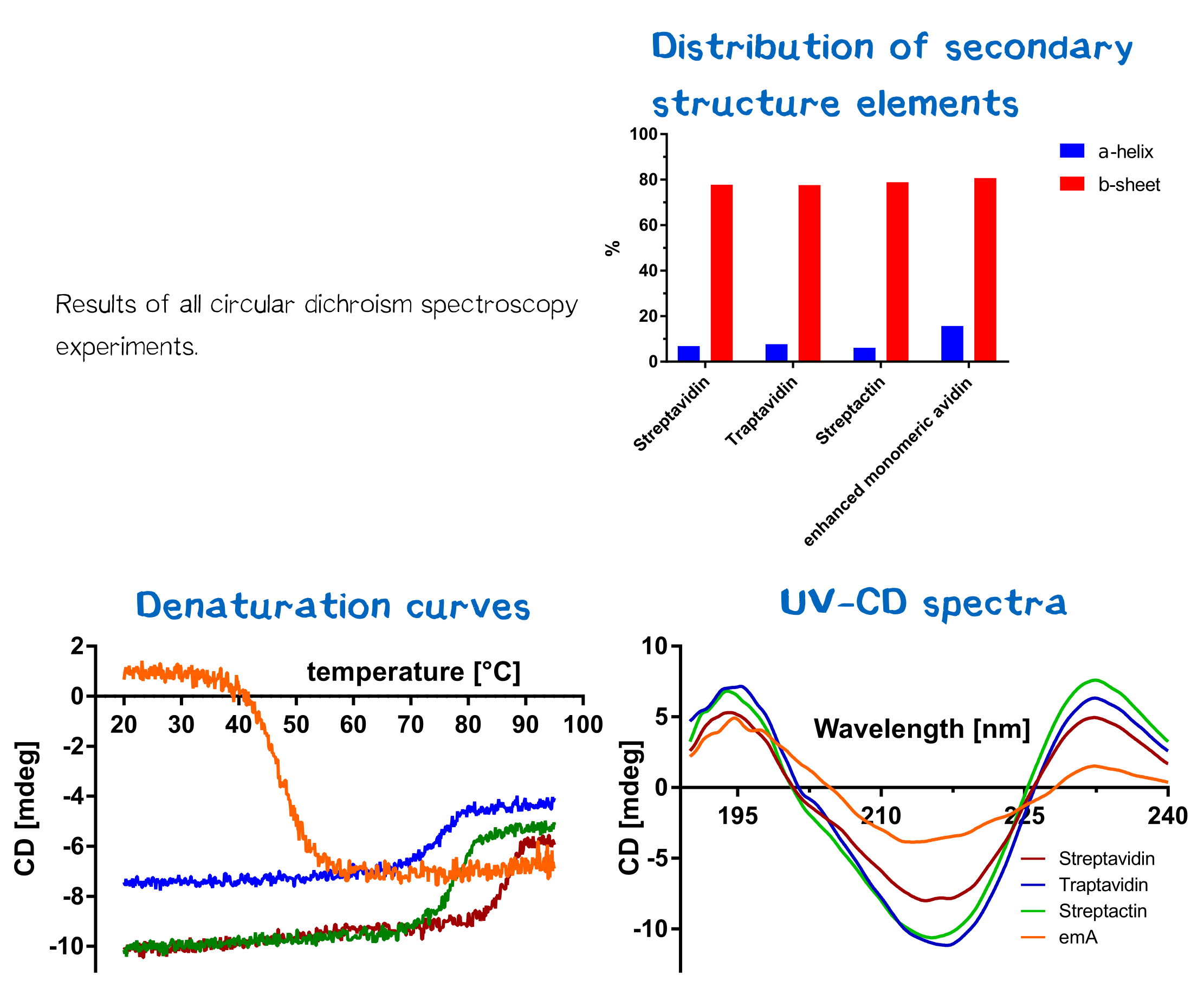
Analysis of secondary structure and thermal stability
De- and renaturation curves as well as UV-CD-spectra were measured and can be seen in figure 14. All cd spectra were evaluated by Spectra Manager Software and results can be seen in figure 14. Also the temperature where 50 % of the protein were unfolded were calculated (Tm. Therefore the data for each protein were entered into a GraphPad Prism sheet and calculated respectively. Results can be seen in table 1. As you can clearly see, Streptavidin and Streptacin are much more stable than emA but less stable than Traptvidin.
Conjugation of proteins with biotin-NHS
References
- ↑ Determined via http://www.expasy.org/
- ↑ Chivers, C. E., Koner, A. L., Lowe, E. D., & Howarth, M. (2011). How the biotin–streptavidin interaction was made even stronger: investigation via crystallography and a chimaeric tetramer. Biochemical Journal, 435(1), 55-63.
- ↑ Weber, P. C., Ohlendorf, D. H., Wendoloski, J. J., & Salemme, F. R. (1989). Structural origins of high-affinity biotin binding to streptavidin. Science, 243(4887), 85.
- ↑ Schmidt, T. G., & Skerra, A. (2007). The Strep-tag system for one-step purification and high-affinity detection or capturing of proteins. Nature protocols, 2(6), 1528-1535. ISO 690
- ↑ Weber, P. C., Ohlendorf, D. H., Wendoloski, J. J., & Salemme, F. R. (1989). Structural origins of high-affinity biotin binding to streptavidin. Science, 243(4887), 85.
- ↑ Stayton, P. S., Freitag, S., Klumb, L. A., Chilkoti, A., Chu, V., Penzotti, J. E., ... & Stenkamp, R. E. (1999). Streptavidin–biotin binding energetics. Biomolecular engineering, 16(1), 39-44.
- ↑ Chivers, C. E., Crozat, E., Chu, C., Moy, V. T., Sherratt, D. J., & Howarth, M. (2010). A streptavidin variant with slower biotin dissociation and increased mechanostability. Nature methods, 7(5), 391-393.
- ↑ Schmidt, T. G., & Skerra, A. (2007). The Strep-tag system for one-step purification and high-affinity detection or capturing of proteins. Nature protocols, 2(6), 1528-1535.
- ↑ Lee, J. M., Kim, J. A., Yen, T. C., Lee, I. H., Ahn, B., Lee, Y., ... & Jung, Y. (2016). A Rhizavidin Monomer with Nearly Multimeric Avidin‐Like Binding Stability Against Biotin Conjugates. Angewandte Chemie International Edition, 55(10), 3393-3397.
- ↑ Skerra, A., Gebauer, M., Schönfeld, D., Schmelz, E. (2007). Skript zu "Fortgeschrittenenpraktikum Proteinchemie"
- ↑ Schlapschy, M., Binder, U., Börger, C., Theobald, I., Wachinger, K., Kisling, S., ... & Skerra, A. (2013). PASylation: a biological alternative to PEGylation for extending the plasma half-life of pharmaceutically active proteins. Protein Engineering Design and Selection, 26(8), 489-501.
- ↑ Majorek, K. A., Porebski, P. J., Dayal, A., Zimmerman, M. D., Jablonska, K., Stewart, A. J., ... & Minor, W. (2012). Structural and immunologic characterization of bovine, horse, and rabbit serum albumins. Molecular immunology, 52(3), 174-182.
- ↑ Cinelli, R. A., Ferrari, A., Pellegrini, V., Tyagi, M., Giacca, M., & Beltram, F. (2000). The Enhanced Green Fluorescent Protein as a Tool for the Analysis of Protein Dynamics and Localization: Local Fluorescence Study at the Single‐molecule Level. Photochemistry and photobiology, 71(6), 771-776.